Drosophila melanogaster is one of the most important model organisms and provides a powerful genetic tool for studies on developmental biology. Drosophila has been successfully used for performing many in vivo studies on molecular functions. Therefore, we established a heritable and inducible RNAi knockdown system in Drosophila for functional analysis of glycans using the GAL4-upstream activating sequence (UAS) system (Fig. 1) (Takemae H. et al. 2003, Nishihara S. et al. 2004, Nishihara S. 2008). |
| Category | Glycogene transgenic animals |
| Protocol Name | Heritable and inducible RNAi knockdown system in Drosophila |
Authors
 |
Nishihara, Shoko
Laboratory of Cell Biology, Department of Bioinformatics, Graduate School of Engineering, Soka University
|
| KeyWords |
|
| Methods |
|
1. |
Heritable and inducible RNAi knockdown system in Drosophila |
| 1) |
Select approximately 500 bp* of the target gene sequence** using dsCheck (http://dscheck.rnai.jp/); this highly sensitive search software avoids off-target effects by using dsRNA-mediated RNAi. |
Comment 1
|

|
| 2) |
Amplify the target gene sequence by polymerase chain reaction (PCR) with a cDNA library derived from Drosophila melanogaster or an expression sequence tag clone (Fig. 2a). The following PCR primer set should be used for this procedure: (1) 5’ primer—AAGGCCTACATGGCCGGACC—together with 21 bp of a target gene sequence that includes an SfiI site and a CpoI site; (2) 3’ primer— AATCTAGAGGTACC—together with 21 bp of a target gene sequence that includes a KpnI site and an XbaI site. |
Comment 0
|

|
| 3) |
The PCR fragment obtained is inserted as an IR sequence into the pUAST-R57 vector (accession number: AB233207) at the abovementioned restriction enzyme sites (Fig. 2b). pUAST-R57 is the modified pUAST vector used for cloning the IRs in the target gene sequence. A fragment comprising exons 5–7 of the fly Ret oncogene is placed between the insertion sites of the IRs. For details of the cloning strategies, please refer to the website http://www.shigen.nig.ac.jp/fly/nigfly/about/aboutRnai.jsp. |
Comment 0
|

|
| 4) |
Inject the constructed vector into Drosophila embryos of the w1118 mutant stock used to establish the UAS-IR transgenic fly lines. Fifty nanograms of IR vector DNA and 50 ng of helper (phs-pai) DNA are transformed into 30 fly eggs*. Refer to the website http://www.shigen.nig.ac.jp/fly/nigfly/about/aboutRnai.jsp for details of the mating scheme (Fig. 3). |
Comment 1
|

|
| 5) |
Mate flies from the UAS-IR fly line with flies from the appropriate GAL4 driver fly lines. Grow the F1 progeny at 25°C or 28°C* and observe their phenotypes. |
Comment 1
|
|
|
| Discussion | Previously, more than 168 human glycan-related transferases (GRTs), including glycosyltransferases and sulfotransferases, have been cloned and their activities identified. Using these human GRT sequences, we performed a Basic Local Alignment Search Tool (BLAST) search of the Drosophila genome database and identified 91 putative GRTs (Nishihara S. 2008). RNAi knockdown of these GRTs can clarify the functions of glycans.
A schema of the heritable and inducible RNAi knockdown system is shown in Fig. 1 (Takemae H. et al. 2003, Nishihara S. et al. 2004, Nishihara S. 2008). We used 2 transgenic fly lines—the GAL4 driver and UAS-inverted repeat (UAS-IR). The GAL4 driver fly has a transgene that contains the yeast transcription factor GAL4; GAL4 expression is controlled by a tissue-specific promoter. The UAS-IR fly has a transgene containing an IR of the target gene that is ligated to the UAS, a target of GAL4. In the F1 generation of these flies, the double-stranded RNA (dsRNA) expression of the target gene is controlled by a tissue-specific promoter, resulting in gene silencing.
We have used this system for 4 glycosyltransferases—protein O-mannosyltransferase 1; protein O-mannosyltransferase 2 (Ichimiya T. et al. 2004); proteoglycan β-1,4-galactosyltransferase I (Takemae H. et al. 2003); and proteoglycan β-1,3-galactosyltransferase II (Ueyama M. et al. 2008)—and for two 3’-phosphoadenosine 5’-phosphosulfate (PAPS) transporters—sll/dPAPST1(Kamiyama et al. 2003) and dPAPST2 (Goda E. et al. 2006). The results clearly indicate that RNAi specifically reduces the expression of target genes in this system and that this reduction induces the development of abnormal phenotypes in flies. |
| Figure & Legends |
Figure & Legends 
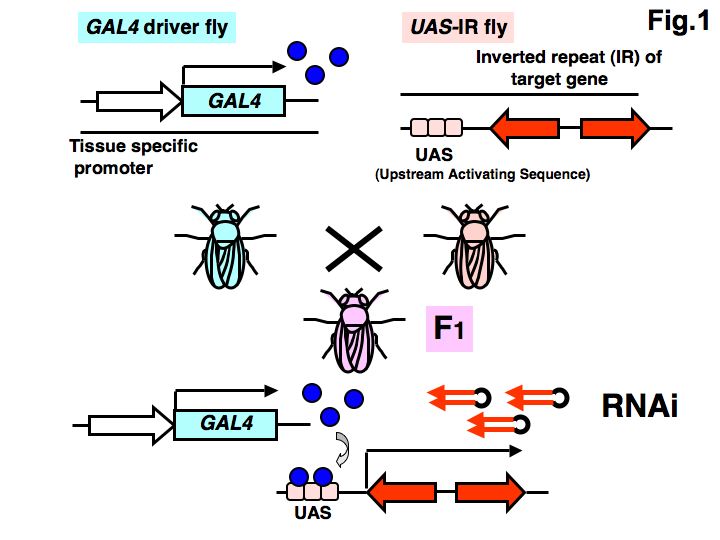
Fig. 1. Schema of the heritable and inducible RNAi system
Two transgenic fly lines—GAL4 driver and UAS-IR—are used in this heritable and inducible RNAi system. The GAL4 driver fly has a transgene containing the yeast transcriptional factor GAL4; GAL4 expression is controlled by a tissue-specific promoter. The UAS-IR fly has a transgene containing an inverted repeat (IR) of the target gene that is ligated to the UAS, a target of GAL4. In the F1 generation of these flies, the dsRNA of the target gene is expressed in a tissue-specific manner and induces gene silencing.
This figure was obtained and modified from "Experimental Glycoscience -Glycobiology" edited by Taniguchi N. et al. Springer Japan KK. 2008, pp.285-289 (Nishihara S. Part2: Section XIV).


Fig. 2. pUAST-R57 vector construction and cloning strategy
pUAST-R57 is the modified pUAST vector used for cloning the IRs of the target gene sequence. A fragment comprising exons 5–7 of the fly Ret oncogene is placed between the insertion sites of the IRs. (a) PCR primer set and cloning strategy, (b) pUAST-R57 vector construction.
This figure was obtained and modified from National Institute of Genetics Fly Stock Center (website: http://www.shigen.nig.ac.jp/fly/nigfly/about/aboutRnai.jsp), Method "Vector construction and cloning strategies (Fig.1)".


Fig. 3. Mating scheme for establishment of UAS-IR transgenic fly lines
This figure was obtained and modified from National Institute of Genetics Fly Stock Center (website: http://www.shigen.nig.ac.jp/fly/nigfly/about/aboutRnai.jsp), Method "Injection and line establishment (Fig. 2)". |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2014-02-26 17:22:52 |
- Takemae, H., Ueda, R., Ohkubo, R., Nakato, H., Izumi, S., Saigo, K., Nishihara, S. (2003) Proteoglycan UDP-galactose: β-xylose β1,4galactosyltransferase I is essential for viability in Drosophila melanogaster. J. Biol. Chem, 278, 15571-15578. [PMID : 12590131]
- Kamiyama, S., Suda, T., Ueda, R., Suzuki, M., Okubo, R., Kikuchi, N., Chiba, Y., Goto, S., Toyoda, H., Saigo, K., Watanabe, M., Narimatsu, H., Jigami, Y., and Nishihara, S. (2003) Molecular cloning and identification of 3'-phosphoadenosine 5'-phosphosulfate transporter. J. Biol. Chem. 278, 25958-25963. [PMID : 12716889]
- Ichimiya, T., Manya, H., Ohmae, Y., Yoshida, H., Takahashi, K., Ueda, R., Endo, T., and Nishihara, S. (2004) The twisted abdomen phenotype of Drosophila POMT1 and POMT2 mutants coincides with their heterophilic protein O-mannosyltransferase activity. J. Biol. Chem. 279, 42638-42647. [PMID : 15271988]
- Nishihara, S., Ueda, R., Goto, S., Toyoda, H., Ishida, H., and Nakamura, M. (2004) Approach for functional analysis of glycan using RNA interference. Glycoconj J. 21, 63-68. [PMID : 15467401]
- Goda, E., Kamiyama, S., Uno, T., Yoshida, H., Ueyama, M., Kinoshita-Toyoda, A., Toyoda, H., Ueda, R., and Nishihara, S. (2006) Identification and characterization of a novel Drosophila 3'-phosphoadenosine 5'-phosphosulfate transporter. J. Biol. Chem. 281, 28508-28517. [PMID : 16873373]
- Nishihara, S. (2007) Drosophila development, RNAi, and glycobiology. Comprehensive Glycoscience - From Chemistry to Systems Biology, edited by Johannis P Kamerling et al. Elsevier. 4.05, 49-79. [PMID : 16610859]
- Ueyama, M., Takemae, H., Ohmae, Y., Yoshida, H., Toyoda, H., Ueda, R., and Nishihara, S. (2008) Functional analysis of proteoglycan galactosyltransferase II RNAi mutant flies. J. Biol. Chem. 283, 6076-6084. [PMID : 18165227]
- Nishihara S (2008) Functional analysis of sugar chains using a genome-wide RNAi system in Drosophila. Experimental Glycoscience - Glycobiology, edited by Naoyuki Taniguchi et al., Springer, Part2: Section XIV, 285-289. [PMID : unknown]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Nishihara, Shoko,
(2014). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.1,5,2024 .
How to Cite this Work in Website:
Nishihara, Shoko,
(2014).
Heritable and inducible RNAi knockdown system in Drosophila.
Retrieved 1,5,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t32.
html source
Nishihara, Shoko,
(2014).
<b>Heritable and inducible RNAi knockdown system in <em>Drosophila</em></b>.
Retrieved 5 1,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t32" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t32</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|