Keratan sulfate is one of the major glycosaminoglycans and consists of poly-N-acetyllactosamine backbone structure [(Galβ1-4GlcNAcβ1-3)n] with modification of sulfates at the 6-OH of Gal and GlcNAc residues. Based on the substrate specificities of glycosyl/sulfotransferases involved in keratan sulfate biosynthesis, keratan sulfate can be elongated as shown in Fig. 1.
- Non-reducing terminal GlcNAc residues on specific branches of N-linked/O-linked glycan chains are 6-O-sulfated by GlcNAc 6-O-sulfotransferase-5 (GlcNAc6ST5, also known as CGn6ST, GST4β, and CHST6).
- GlcNAc 6-O-sulfate residues are next galactosylated by β1,4-galactosyltransferase-IV (β4GalT-IV), which is only the β4GalT specific for GlcNAc 6-O-sulfate.
- 6-O-Sulfated LacNAc, Galβ1-4(SO3--6)GlcNAc, is elongated by β1,3-N-acetylglucosaminyltransferase-7 (β3Gn-T7), which is specific for the sulfated moiety.
- Repeating 1, 2, and 3, resulting in the formation of [Galβ1-4(SO3--6)GlcNAcβ1-3]n.
- Gal 6-O-sulfotransferase (Gal6ST, also known as KSGal6ST, GST-1, and CHST-1) catalyzes 6-O-sulfation of both the non-reducing terminal and internal Gal residues.
As described above, two sulfotransferases, GlcNAc6ST5 and Gal6ST, are involved in the biosynthesis of keratan sulfate. The enzymatic activities of these enzymes can be measured using with a donor substrate, adenosine 3’-phosphate 5’-phospho[35S]sulfate ([35S]PAPS) and acceptor substrates bearing appropriate non-reducing terminal sugar residues. |
| Category | Glycosyltransferases & related proteins |
| Protocol Name | Enzyme assay of sulfotransferases for keratan sulfate |
Authors
 |
Seko, Akira
Ito Glycotrilogy Project, Japan Science and Technology Agency (JST)
Yamashita, Katsuko
*
Innovative Research Initiatives, Tokyo Institute of Technology
*To whom correspondence should be addressed.
|
| KeyWords |
|
Reagents
 |
| ● |
Adenosine 3’-phosphate 5’-phospho[35S]sulfate (PerkinElmer, Waltham, MA) |
| ● |
Keratan sulfate (Seikagaku Corp., Tokyo, Japan) |
|
Instruments
 |
| ● |
High-voltage paper electrophoresis unit (Advantec Co., Ltd. Ehime, Japan) |
| ● |
Radiochromatogram scanner (RITA, Raytest, Straubenhardt, Germany) |
|
| Methods |
|
1. |
Preparation of an intermediate analogue of keratan sulfate, GlcNAcβ1-3Galβ1-4(SO3--6)GlcNAcβ1-3Galβ1-4(SO3--6)GlcNAc (GlcNAcβ1-3L2L2) |
| 1) |
0.1 mL of the reaction mixture containing 50 mM HEPES-NaOH (pH 7.2), 10 mM MnCl2, 0.05%(v/v) Triton X-100, 50 μg/mL protamine chloride, 1 mM L2L2 (Ref. 3), 0.1 M GlcNAc, 1.25 mM UDP-GlcNAc, and recombinant β3Gn-T7 (5 nmol/h. The activity is assayed using 0.5 mM L2L2 and 0.5 mM UDP-GlcNAc), is incubated at 37ºC for 1 day. |
Comment 1
|

|
| 2) |
The mixture is applied to Sephadex G-50 gel filtration (1.3 × 68 cm, equilibrated and eluted with 0.1 M NaCl). |
Comment 0
|

|
| 3) |
The hexose-positive fractions (phenol-sulfuric acid method) are applied to RCA-I-agarose (J-OIL MILLS, Inc., Tokyo, Japan) lectin chromatography to remove residual L2L2. |
Comment 0
|

|
| 4) |
The pass-through fractions are desalted by Sephadex G-25 gel filtration (1.3 × 68 cm, equilibrated and eluted with EtOH/water 1:19). Finally, 27 nmol of GlcNAcβ1-3L2L2 is obtained. |
Comment 0
|
|
|
|
2. |
|
| 1) |
Twenty μL of the reaction mixture containing 50 mM sodium cacodylate (pH 6.8), 10 mM MnCl2, 0.1 % (w/v) digitonin (if enzymes to be analyzed would be membrane fractions), 50 μg/mL protamine chloride, 2 mM dithiothreitol, 0.1 M NaF, 2 mM ATP-Na2, 6.5 μM [35S]PAPS (4.9 × 105 dpm), 0.1 mM GlcNAcβ1-3L2L2, and the enzyme fractions, is incubated at 37ºC for 1 h. |
Comment 0
|

|
| 2) |
Add 0.5 mL of 0.01 N HCl and heat at 100ºC for 10 min to destroy residual [35S]PAPS. After cooling, the mixture is neutralized and concentrated by vacuum evaporator. |
Comment 0
|

|
| 3) |
Spot the mixture and bromophenol blue as a marker, on a Whatman No. 1 paper (46 × 57 cm). |
Comment 0
|

|
| 4) |
Wet the paper with pyridine/acetic acid/water=3:1:387 (pH 5.4). |
Comment 0
|

|
| 5) |
Set the paper into the high-voltage paper electrophoresis unit and perform electrophoresis at 4,000 V with the same solvent as 4), until the marker moves 10 cm from the origin. |
Comment 0
|

|
| 6) |
Dry the paper in the draft and measure the radioactivity using with a radiochromatogram scanner. Generally, [35S]-labeled reaction product moves near the marker, while free sulfate moves about 2.5-fold forward from the marker. |
Comment 0
|
|
|
|
3. |
|
| 1) |
Prepare the reaction mixture as above, except for adding 50 mM sodium cacodylate (pH 6.4), 0.1 % (v/v) Triton X-100, and 0.5 mg/mL keratan sulfate in place of sodium cacodylate (pH 6.8), digitonin, and GlcNAcβ1-3L2L2, respectively. |
Comment 0
|

|
| 2) |
Same as above step 2-6). [35S]-labeled reaction products move as smear profile between origin and the marker. |
Comment 0
|
|
|
| Notes |
- GlcNAc6ST5 and Gal6ST belongs to GlcNAc6ST family which consists of 7 members in human, GlcNAc6ST1 to 5, Gal6ST and C6ST-1. If enzyme fractions to be analyzed are crude ones, it is very difficult to determine the individual enzymatic activities of GlcNAc6ST5 and Gal6ST, because some of other members can utilize keratan sulfate or GlcNAcβ1-3L2L2 as donor substrate. For example, not only Gal6ST, but also chondroitin 6-O-sulfotransferase 1 (C6ST-1) can act on keratan sulfate. Similarly, GlcNAcβ1-3L2L2 can be utilized by GlcNAc6ST1, 4, and 5, and Gal6ST. GlcNAc6ST1, 4, and 5 can catalyze sulfation of the 6-OH of non-reducing terminal GlcNAc of GlcNAcβ1-3L2L2, although the ratio of Vmax/Km values of the three enzymes for the substrate is 26:4:100, indicating the preference of GlcNAc6ST5 for the substrate. Gal6ST can catalyze sulfation of internal Gal residues of the substrate.
- Their recombinant enzymes produced in E. coli would be generally inactive, probably by their insolubility or improper folding. Other hosts such as Pichia pastoris or culture cells (CHO, COS, and so on) are recommended.
- There is danger of an electric shock when using high-voltage paper electrophoresis unit. Power source must be turned off when papers would be set into the unit or taken out from it.
|
| Figure & Legends |
Figure & Legends 
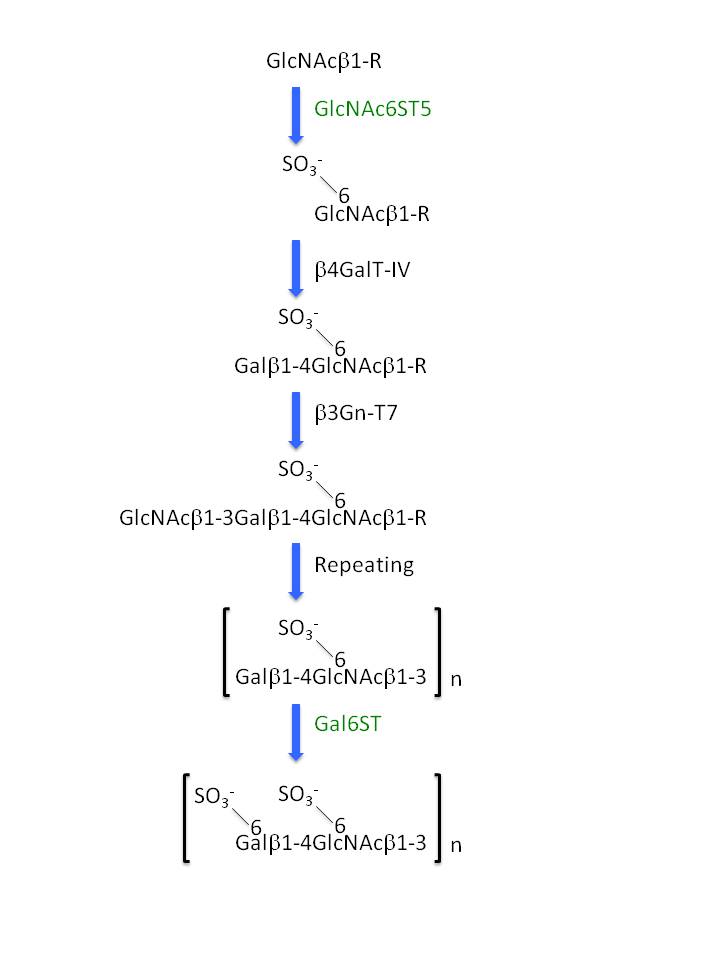
Fig. 1. Biosynthesis of keratan sulfate. See Introduction in details for respective steps. |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2014-05-27 13:14:43 |
- Torii ,T., Fukuta, M., and Habuchi, O. (2000) Sulfation of sialyl N-acetyllactosamine oligosaccharides and fetuin oligosaccharides by keratan sulfate Gal-6-sulfotransferase. Glycobiology 10, 203–211. [PMID : 10642612]
- Akama, T.O., Misra, A.K., Hindsgaul, O., and Fukuda M.N. (2002) Enzymatic synthesis in vitro of the disulfated disaccharide unit of corneal keratin sulfate. J. Biol. Chem. 277, 42505–42513. [PMID : 12218059]
- Brown, G.M., Huckerby, T.N., and Nieduszynski I.A. (1994) Oligosaccharides derived by keratanase II digestion of bovine articular cartilage keratin sulphates. Eur. J. Biochem. 224, 281–308. [PMID : 7925342]
- Seko, A., Nagata, K., Yonezawa, S., and Yamashita, K. (2002) Ectopic expression of a GlcNAc 6-O-sulfotransferase, GlcNAc6ST-2, in colonic mucinous adenocarcinoma. Glycobiology 12, 379–388. [PMID : 12107080]
- Seko, A., and Yamashita, K. (2004) b1,3-N-Acetylglucosaminyltransferase-7 (b3Gn-T7) acts efficiently on keratan sulfate-related glycans. FEBS Lett. 556, 216–220. [PMID : 14706853]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Seko, Akira,
Yamashita, Katsuko,
(2014). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.20,4,2024 .
How to Cite this Work in Website:
Seko, Akira,
Yamashita, Katsuko,
(2014).
Enzyme assay of sulfotransferases for keratan sulfate.
Retrieved 20,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t88.
html source
Seko, Akira,
Yamashita, Katsuko,
(2014).
<b>Enzyme assay of sulfotransferases for keratan sulfate</b>.
Retrieved 4 20,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t88" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t88</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|