EXT1 and EXT2 genes encode heparan sulfate (HS) co-polymerases, and the genetic defects of which result in hereditary multiple exostoses in humans. Both EXT1 and EXT2 exhibit GlcNAc transferase (GlcNAcT) and GlcA transferase (GlcAT) activities required for the HS synthesis (Fig. 1). EXT1 and EXT2 form a stable complex and the complex is thought to be the biologically relevant form of enzymes. In addition, three highly homologous EXT-like genes, EXTL1-EXTL3, have been cloned (Fig. 1). Both EXTL1 and EXTL3 have been shown to exhibit GlcNAcT activities likely involved in the biosynthesis of HS (Fig. 1). EXTL1 exerts only GlcNAcT-II activity, which may be involved in HS chain elongation, whereas EXTL3 possesses activity transferring the first GlcNAc residue to tetrasaccharide-linkage region (so-called GlcNAcT-I activity) in addition to GlcNAcT-II activities. EXTL2 exhibits the GlcNAcT-I activities involved in the initiation of HS biosynthesis (Fig. 1). Although all EXTL proteins show glycosyltrasferase activities associated with HS biosynthesis, their roles in the HS biosynthesis remain unclear. |
| Category | Glycosyltransferases & related proteins |
| Protocol Name | Enzyme assay of GAG glycosyltransferases for heparan sulfate |
Authors
 |
Nadanaka, Satomi
Department of Biochemistry, Kobe Pharmaceutical University
Kitagawa, Hiroshi
*
Department of Biochemistry, Kobe Pharmaceutical University
*To whom correspondence should be addressed.
|
| KeyWords |
|
Reagents
 |
| ● |
GlcNAcT-I activities
Substrate: GlcAβ1-3Galβ1-O-C2H4NH-benzyloxycarbonyl (chemically-synthesized linkage region analog)*
Donor: UDP-[3H]GlcNAc (60 Ci/mmol) (PerkinElmer, Waltham, MA)
* Acceptor substrates for GlcNAcT-I, GlcAβ1-3Galβ1-O-C2H4NH-benzyloxycarbonyl, were kindly provided from Prof. Jun-ichi Tamura (Tottori University). |
| ● |
GlcNAcT-II activities
Substrate: N-Acetylheparosan oligosaccharides derived from the capsular polysaccharide of Escherichia coli K5,
GlcAβ1-4GlcNAcα1-(4GlcAβ1-4GlcNAcα1)n with the non-reducing terminal GlcA
Donor: UDP-[3H]GlcNAc (60 Ci/mmol) (PerkinElmer) |
| ● |
HS-GlcAT-II activities
Substrate: N-Acetylheparosan oligosaccharides derived from the capsular polysaccharide of Escherichia coli K5,
GlcNAcα1-(4GlcAβ1-4GlcNAcα1)n with the non-reducing terminal GlcNAc
Donor: UDP-[U - 14C]GlcA (285.2 mCi/mmol) (PerkinElmer) |
| ● |
Polymerization activities
Substrate: GlcAβ1-3Galβ1-O-C2H4NH-benzyloxycarbonyl (chemically-synthesized linkage region analog)* or
GlcAβ1-3Galβ1-3Galβ1-4Xylβ1-O-Ser-Gly-Trp-Pro-Asp-Gly (chemically-synthesized) 1)
Donor: UDP-[3H]GlcNAc (60 Ci/mmol) and UDP-[U - 14C]GlcA (285.2 mCi/mmol) (PerkinElmer)
* Acceptor substrates for GlcNAcT-I, GlcAβ1-3Galβ1-O-C2H4NH-benzyloxycarbonyl, were kindly provided from Prof. Jun-ichi Tamura (Tottori University). |
|
Instruments
 |
| ● |
GlcNAcT-I activities
Nova-Pak C18 column (3.9 × 150 mm) (Waters Corp., Milford, MA)
HPLC
2300TR liquid scintillation counter (PerkinElmer) |
| ● |
GlcNAcT-II activities and HS-GlcAT-II activities
Syringe column (TERUMO SS-01T) packed with Superdex G-25 superfine (GE Healthcare, Little Chalfont, UK) 2)
2300TR liquid scintillation counter (PerkinElmer) |
| ● |
Polymerization activities
Superdex Peptide HR 10/30 or Superdex 200 10/300 GL (GE Healthcare)
FPLC
2300TR liquid scintillation counter (PerkinElmer) |
|
| Methods |
|
1. |
Enzyme assay of GAG glycosyltransferases for heparan sulfate |
| 1) |
The following reaction mixtures of a total volume of 20 μL are prepared.
GlcNAcT-I activities
10 μL of enzyme source (see Comment *)
100 mM MES-NaOH (pH 6.5)
10 mM MnCl2
1 mM ATP-2Na
GlcAβ1-3Galβ1-O-C2H4NH-benzyloxycarbonyl (250 nmol)
250 μM UDP-[3H]GlcNAc (approx. 1.1 × 106 dpm)
GlcNAcT-II activities
10 μL of enzyme source
100 mM MES-NaOH (pH 6.5)
10 mM MnCl2
1 mM ATP-2Na
GlcAβ1-4GlcNAcα1-(4GlcAβ1-4GlcNAcα1)n (20 μg) (see Comment **)
250 μM UDP-[3H]GlcNAc (approx. 1.1 × 106 dpm)
HS-GlcAT-II activities
10 μL of enzyme source
100 mM MES-NaOH (pH 6.5)
10 mM MnCl2
10 mM MgCl2
5 mM CaCl2
171 μM ATP-2Na
GlcNAcα1-(4GlcAβ1-4GlcNAcα1)n (20 μg) (see Comment **)
250 μM UDP-[14C]GlcA (approx. 1.4 × 105 dpm)
Polymerization activities
10 μL of enzyme source
100 mM MES-NaOH (pH 6.5)
10 mM MnCl2
1 mM ATP-2Na
GlcAβ1-3Galβ1-O-C2H4NH-benzyloxycarbonyl (100 nmol)
250 μM UDP-[3H]GlcNAc (approx. 5.5 × 105 dpm)
250 μM UDP-GlcA |
Comment 1
|

|
| 2) |
Reaction mixtures of GlcNAcT-I, GlcNAcT-II, and HS-GlcAT-II are incubated at 37˚C for 4 h.
Polymerization reaction is performed at 37˚C overnight. |
Comment 0
|

|
| 3) |
Each enzyme activity is measured according to the following method.
GlcNAcT-I activities (Fig. 2) 3)
GlcNAcT-I reaction products
↓ HPLC analysis on a Nova-Pak C18 column.
↓ the effluent fractions are analyzed for radioactivity.
GlcNAcT-II and HS-GlcAT-II activities (Fig. 3)
i) Preparation of syringe columns
Syringe columns (TERUMO SS-01T)
↓ pack with Sephadex G-25 (superfine) equilibrated with 0.2 M NH4HCO3.
↓ centrifuge at 800 rpm for 5 min.
↓ centrifuge again at 2,000 rpm for 5 min.
ii) Assay
Samples (20 μL)
↓ add 30 μL of 0.2 M NH4HCO3.
Samples (50 μL)
↓ apply to the packed syringe column.
↓ centrifuge at 2,000 rpm for 5 min.
↓ add 50 μL of 0.2 M NH4HCO3 to the top of the syringe columns.
↓ centrifuge at 2,000 rpm for 5 min.
Eluates
↓ measure by scintillation counting.
Polymerization activities (Fig. 4)
The reaction products
↓ analyze using a Superdex Peptide HR 10/30 column (see Comment *).
↓ measure radioactivity using a liquid scintillation counter. |
Comment 1
|

|
| 4) |
Each reaction product is confirmed.
GlcNAcT-I-reaction products 3)
GlcNAcT-I reaction products (Fig. 2)
↓ pool and evaporate.
GlcNAcT-I reaction products (about 36 pmol)
↓ digest with 10 mIU of heparitinase I (Seikagaku Corp., Tokyo, Japan) in a total volume of
50 μL of 20 mM sodium acetate buffer (pH 7.0) containing 2 mM calcium
acetate at 37˚C overnight.
↓ analyze digested or undigested samples on a Nova-Pak C18 column.
The radioactivity of the GlcNAcT-I-reaction products is released by digestion
with heparitinase I and eluted in the flow-through fraction as free [3H]GlcNAc.
GlcNAcT-II-reaction products 3)
GlcNAcT-II reaction products
↓ pool and evaporate.
GlcNAcT-II reaction products (about 36 pmol)
↓ digest with 10 mIU of heparitinase I (Seikagaku Corp.) in a total volume of
100 μL of 20 mM sodium acetate buffer (pH 7.0) containing 2 mM calcium
acetate at 37˚C overnight.
↓ analyze by a Superdex peptide HR 10/30 column.
The radioactivity peak of the GlcNAcT-II-reaction products eluted near the void volume
is released by digestion with heparitinase I and shifted to the free [3H]GlcNAc position.
Polymerization activities 1)
Polymerized products (Fig. 4)
↓ pool and evaporate.
Polymerized products (about 70 pmol as disaccharide)
↓ digest with 6 mIU of heparitinase I (Seikagaku Corp.) in a total volume of 100 μL of
20 mM sodium acetate buffer (pH 7.0) containing 2 mM calcium acetate at 37˚C overnight.
↓ analyze using a Superdex peptide HR 10/30 column.
The radioactivity peak of the polymerized products eluted near the void volume is
degraded by digestion with heparitinase I and shifted to the elution position of the
unsaturated disaccharide. |
Comment 0
|
|
|
| Figure & Legends |
Figure & Legends 
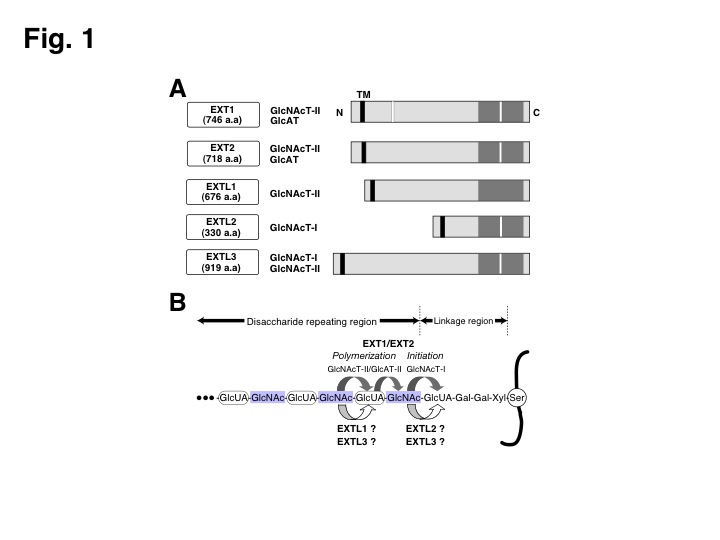
Fig. 1. The five cloned members of the EXT gene family.
(A) Schematic structures of five EXT gene family are shown. Highly conserved regions are indicated by gray bars. White bars show DXD motif. a.a., amino acid; TM, transmembrane domain. In addition, glycosyltransferase activities detected in EXT members are summarized. (B) The biosynthesis of HS in mammals. Mammalian HS is biosynthesized by EXT1/EXT2 polymerase complex. The significance of EXTL1, EXTL2, and EXTL3 in the biosynthesis of HS remains unclear.

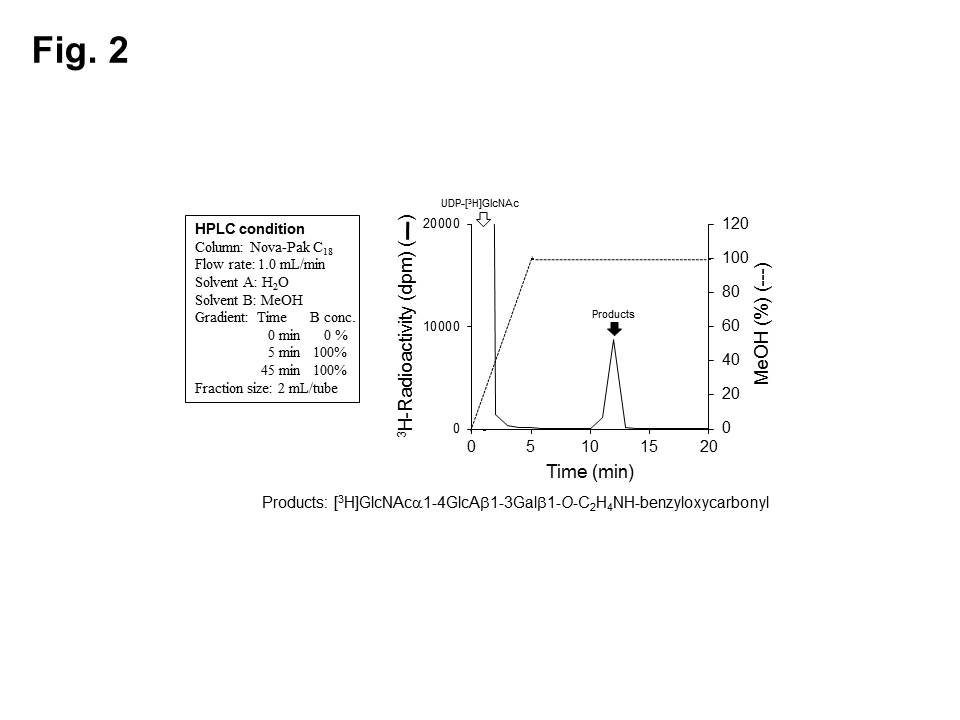
Fig. 2. Measurement of GlcNAcT-I activities.
GlcNAcT-I-reaction products were subjected to HPLC analysis using a Nova-Pak C18 column. The separated fractions (2 mL each) were measured for radioactivities. White arrow, UDP-[3H]GlcNAc; Black arrow, [3H]GlcNAca1-4GlcAb1-3Galb1-O-C2H4NH-benzyloxycarbonyl.

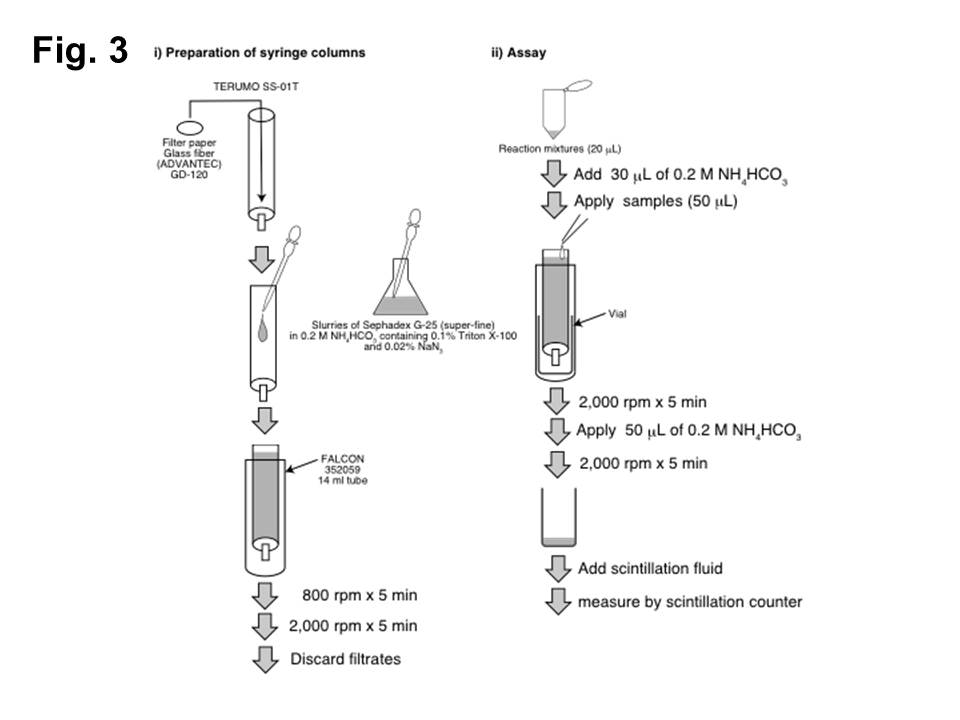
Fig. 3. GlcNAcT-II and HS-GlcAT-II assays using a syringe column.

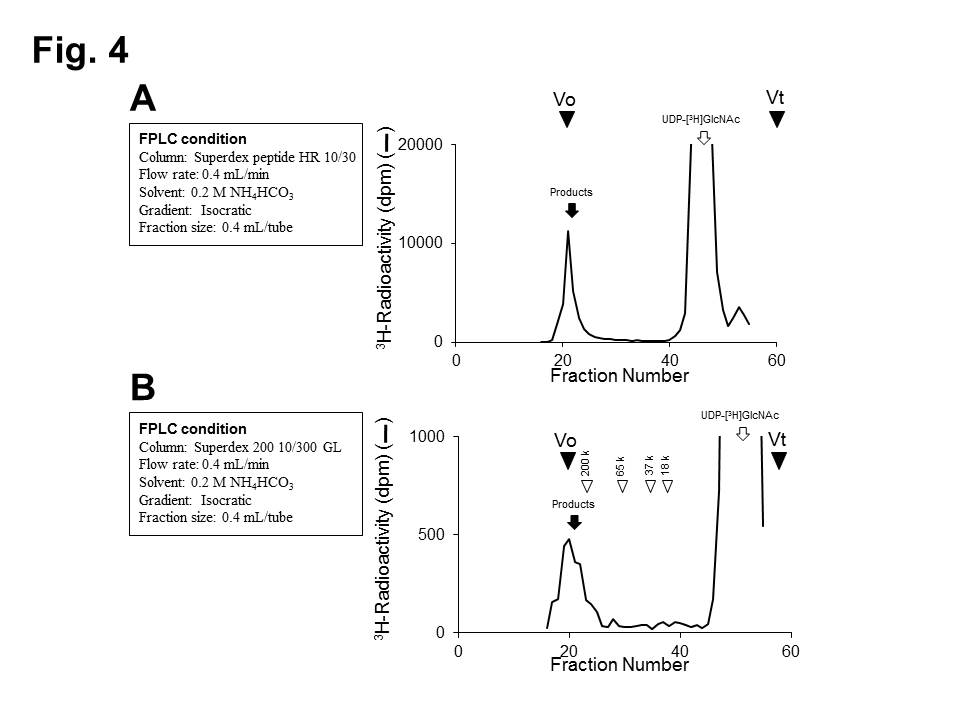
Fig. 4. Analysis of polymerized HS chains.
(A) The polymerization reaction products were analyzed on a Superdex peptide column .
(B) Molecular size of HS chains was determined by gel filtration chromatography on a Superdex 200 column. White arrowheads indicate the elution positions of commercial dextrans of known molecular weights. |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2014-09-11 16:20:53 |
- Kim, B.T., Kitagawa, H., Tanaka, J., Tamura, J., and Sugahara, K. (2003) In vitro heparan sulfate polymerization: crucial roles of core protein moieties of primer substrates in addition to the EXT1-EXT2 interaction. J Biol Chem. 278, 41618–41623 [PMID : 12907685]
- Kitagawa, H., Tsuchida, K., Ujikawa, M., and Sugahara, K. (1995) Detection and characterization of UDP-GalNAc: chondroigin N-acetylgalactosaminyltransferase in bovine serum using a simple assay method. J Biochem. 117, 1083–1087 [PMID : 8586623]
- Kim, B.T., Kitagawa, H., Tamura, J., Saito, T., Kusche-Gullberg, M., Lindahl, U., and Sugahara, K. (2001) Human tumor suppressor EXT gene family members EXTL1 and EXTL3 encode alpha 1,4-N-acetylglucosaminyltransferases that likely are involved in heparan sulfate/heparin biosynthesis. Proc Natl Acad Sci U S A. 98, 7176–7181 [PMID : 11390981]
- Busse, M., Feta, A., Presto, J., Wilen, M., Gronning, M., Kjellen, L., and Kusche-Gullberg, M. (2007) Contribution of EXT1, EXT2, and EXTL3 to heparan sulfate chain elongation. J Biol Chem. 282, 32802–32810 [PMID : 17761672]
- Lidholt, K., Weinke, J.L., Kiser, C.S., Lugemwa, F.N., Bame, K.J., Cheifetz, S., Massague, J., Lindahl, U., and Esko, J.D. (1992) A single mutation affects both N-acetylglusosaminyltransferase and glucuronosyltransferase activities in a Chinese hamster ovary cell mutant defective in heparan sulfate biosynthesis. Proc Natl Acad Sci U S A. 89, 2267–2271 [PMID : 1532254]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Nadanaka, Satomi,
Kitagawa, Hiroshi,
(2014). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.25,4,2024 .
How to Cite this Work in Website:
Nadanaka, Satomi,
Kitagawa, Hiroshi,
(2014).
Enzyme assay of GAG glycosyltransferases for heparan sulfate.
Retrieved 25,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t83.
html source
Nadanaka, Satomi,
Kitagawa, Hiroshi,
(2014).
<b>Enzyme assay of GAG glycosyltransferases for heparan sulfate</b>.
Retrieved 4 25,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t83" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t83</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|