Many eukaryotic cell surface proteins with various functions are anchored to the membrane via glycosylphosphatidylinositol (GPI). GPI-anchored proteins are released from the plasma membrane by bacterial phosphatidylinositol-specific phospholipase C (PI-PLC). Therefore, PI-PLC is used as a tool to determine whether proteins of interest are GPI-anchored. Here, we describe a protocol for PI-PLC treatment of GPI-anchored proteins on intact cells for detection by flow cytometry. |
| Category | GPI anchored proteins |
| Protocol Name | Assay of genes involved in GPI biosynthesis in ER ~PI-PLC treatment of GPI-anchored proteins on the cell surface of intact cells |
Authors
 |
Fujita, Morihisa
Research Institute for Microbial Diseases, Osaka University
Maeda, Yusuke
Research Institute for Microbial Diseases, Osaka University
Kinoshita, Taroh
*
Research Institute for Microbial Diseases, Osaka University
*To whom correspondence should be addressed.
|
| KeyWords |
|
Reagents
 |
| ● |
Phosphate-buffered saline (PBS) |
| ● |
Phosphatidylinositol-specific phospholipase C (PI-PLC) |
| ● |
|
| ● |
OPTI-MEM (Invitrogen-Life Technologies, Carlsbad, CA) |
|
Instruments
 |
| ● |
Flow cytometer (example: BD FACSCalibur (BD Biosciences, San Jose, CA)) |
|
| Methods |
|
1. |
PI-PLC treatment of GPI-anchored proteins on the cell surface of intact cells |
| 1) |
Suspend cells (1 × 106) in 1 mL PBS. |
Comment 0
|

|
| 2) |
Centrifuge at 300 × g for 3 min and resuspend pellet in 100 μL OPTI-MEM. |
Comment 0
|

|
| 3) |
Divide suspension into two (50 μL each) and add PI-PLC (1 unit/mL) or 50% glycerol (control). |
Comment 0
|

|
| 5) |
Add 500 μL PBS, centrifuge at 300 × g for 3 min and remove the supernatant. |
Comment 0
|

|
| 6) |
Stain proteins of interest with the antibodies and detect the surface expression by flow cytometry. |
Comment 0
|
|
|
| Notes | Inositol-acylated GPI anchors are not cleaved by PI-PLC. During GPI biosynthesis in the endoplasmic reticulum (ER), an acyl-chain is transferred to the 2-position of inositol on the GPI intermediate. Therefore, GPI intermediates are resistant to PI-PLC. Soon after GPI attachment to proteins, the acyl-chain is usually eliminated by GPI-inositol deacylase PGAP1 in the ER. The deacylated GPI anchors subsequently become sensitive to PI-PLC. Therefore, GPI-anchored proteins on mammalian cells are usually sensitive to PI-PLC. However, GPI-anchored proteins on human and mouse erythrocytes are resistant to PI-PLC treatment because the GPI structures maintain the acyl-chain on the 2-position of the inositol. |
| Figure & Legends |
Figure & Legends 
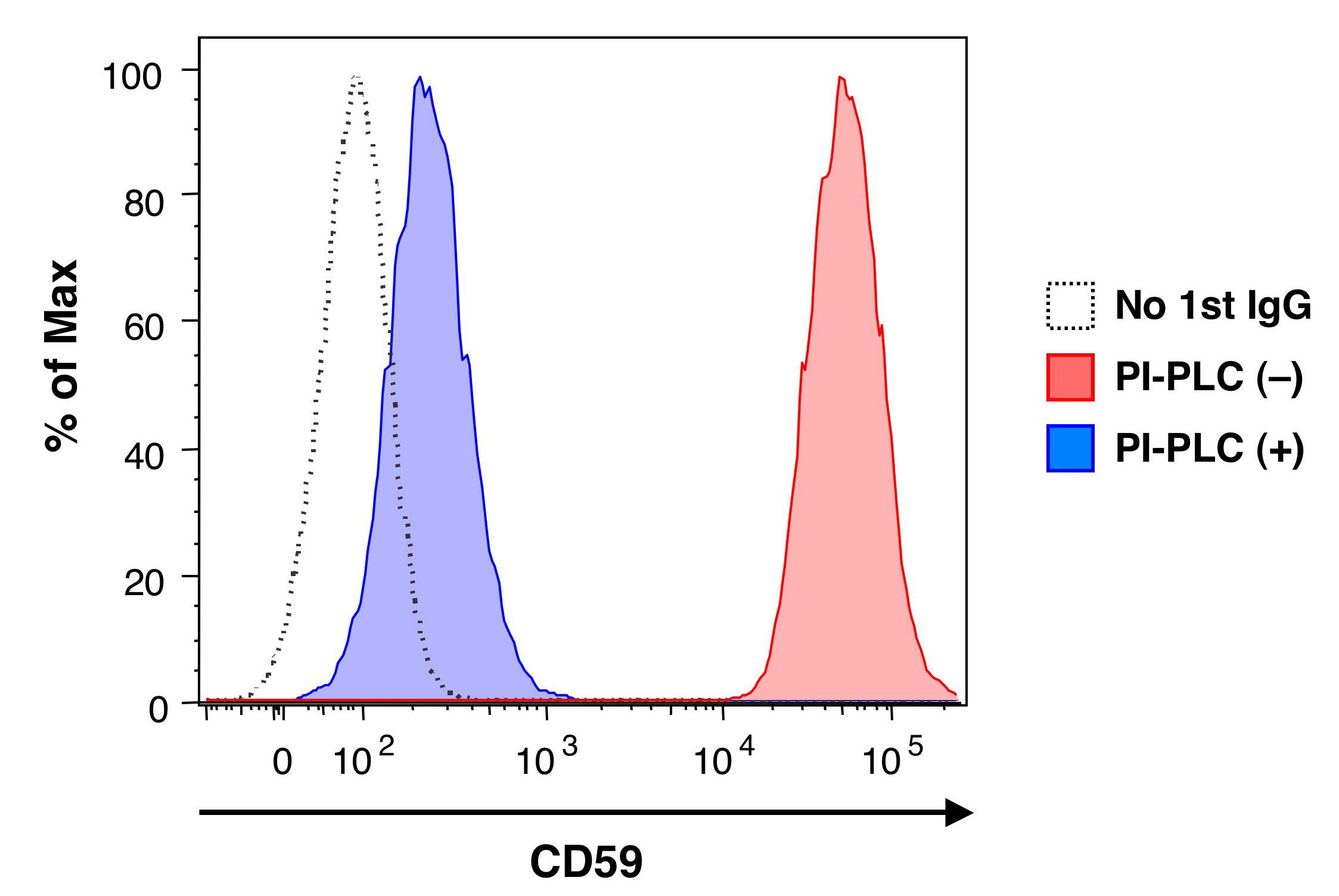
Fig. 1. PI-PLC sensitivity of the cell surface GPI-anchored proteins.
CHO-K1 cells stably expressing human CD59 were treated with (blue) or without (red) PI-PLC. Surface CD59 was stained with anti-CD59 (5H8) followed by phycoerythrin-conjugated anti-mouse IgG, and the cells were analyzed by flow cytometry. Dotted line indicates negative control staining with phycoerythrin-conjugated anti-mouse IgG only. |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2015-01-07 14:18:06 |
- Tanaka, S., Maeda, Y., Tashima, Y., and Kinoshita, T. (2004) Inositol-deacylation of glycosylphosphatidylinositol-anchored proteins is mediated by mammalian PGAP1 and yeast Bst1p. J. Biol. Chem. 279, 14256–14263 [PMID : 14734546]
- Doering, T. L., Englund, P.T., and Hart, G.W. (2001) Detection of Glycophospholipid Anchors on Proteins. Current Protocols in Molecular Biology 17.8.1–17.8.13 [PMID : 18265164]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Fujita, Morihisa,
Maeda, Yusuke,
Kinoshita, Taroh,
(2015). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.27,4,2024 .
How to Cite this Work in Website:
Fujita, Morihisa,
Maeda, Yusuke,
Kinoshita, Taroh,
(2015).
Assay of genes involved in GPI biosynthesis in ER ~PI-PLC treatment of GPI-anchored proteins on the cell surface of intact cells.
Retrieved 27,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t75.
html source
Fujita, Morihisa,
Maeda, Yusuke,
Kinoshita, Taroh,
(2015).
<b>Assay of genes involved in GPI biosynthesis in ER ~PI-PLC treatment of GPI-anchored proteins on the cell surface of intact cells</b>.
Retrieved 4 27,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t75" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t75</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|