Endoglycosidases cleave at defined sites within an oligosaccharide chain of glycoproteins/glycolipids. They are the most easy-to-use enzymes for elucidating the function and structure of oligosaccharides, because they can separate both intact oligosaccharide chains and proteins/lipids from glycoconjugates under mild conditions without causing damage.
Endo-β-N-acetylglucosaminidase (EC 3.2.1.96) catalyzes the hydrolysis of the N, N’-diacetylchitobiose moiety in the core region of asparagine-linked oligosaccharides of various glycoproteins. The enzyme has a characteristic to remain one N-acetyl-D-glucosamine residue on the protein. On the other hand, the deglycosylation method using PNGase F cannot remove oligosaccharides unless the protein is denatured. Thus, only endo-β-N-acetylglucosaminidases can be used for deglycosylation of native glycoproteins.
Endo-β-N-acetylglucosaminidase H (Endoglycosidase H or Endo-H) has high specificity for high-mannose type oligosaccharides and for some hybrid-type oligosaccharides, but not for complex-type oligosaccharides (Fig. 1) 1,2). Endo-H is used for the structural study of the N-linked glycans in glycoproteins 3,4,5). |
| Category | N-Glycans |
| Protocol Name | Endo-β-N-acetylglucosaminidase H digestion (Endo-H) |
Authors
 |
Fujita, Kiyotaka
Faculty of Agriculture, Kagoshima University
Yamamoto, Kenji
*
Research Institute for Bioresources and Biotechnology, Ishikawa Prefectural University
*To whom correspondence should be addressed.
|
| KeyWords |
|
Reagents
 |
| ● |
Endo-β-N-acetylglucosaminidase H (Endoglycosidase H or Endo-H) from Streptomyces plicatus.
- Enzymes purified from culture filtrate are commercially available from Sigma-Aldrich (St. Louis, MO), Seikagaku Biobusiness Co. (Tokyo, Japan), AMS Biotechnology (Witney, UK) and United States Biological (Swampscott, MA).
- Recombinant enzymes (expressed in E. coli) are commercially available from Roche Diagnostics GmbH (Mannheim, Germany), New England Bio Labs Inc. (Billerica, MA), Sigma-Aldrich (St. Louis, MO), QA-Bio (San Mateo, CA), AMS Biotechnology (Witney, UK), Prozyme (San Leandro, CA) and Merck KGaA (Darmstadt, Germany).
|
| ● |
Denaturation solution: 2% w/v sodium lauryl sulfate (SDS), 1 M β-mercaptoethanol (β-ME) |
| ● |
5X Reaction buffer : 250 mM sodium phosphate buffer (pH5.5) |
| ● |
2X SDS-PAGE sample buffer: 0.125 M Tris-HCl buffer (pH 6.8), 10% β-ME, 4% SDS, 10% sucrose, 0.004% Bromophenol blue |
|
Instruments
 |
| ● |
Reaction incubator or water bath (100°C, 37 °C) |
| ● |
|
| ● |
Microcon Ultracel YM-10 (Millipore) |
|
| Methods |
|
1. |
Release of oligosaccharides from glycoproteins by using Endo-H |
| 1) |
Transfer 20 μL of glycoprotein sample (10 μg/μL), 10 μL of 5X Reaction buffer, and 17.5 μL of deionized water into a microtube. |
Comment 1
|

|
| 2) |
Add 2 μL of Endo-H solution to attain a final enzyme concentration of 0.5~10 U /mL. |
Comment 0
|

|
| 4) |
Terminate the reaction by heating at 100°C for 3 min. |
Comment 1
|

|
| 5) |
Separate the oligosaccharides from the deglycosylated protein by ultrafiltration using Microcon Ultracel YM-10 (10-kDa cut-off membrane; Millipore). |
Comment 1
|
|
|
|
2. |
SDS-PAGE analysis of glycoproteins deglycosylated with Endo-H. |
| 1) |
Transfer 20 μL of glycoprotein sample (10 μg/μL), 10 μL of 5X reaction buffer, 2.5 μL of denaturation solution, and 15.5 μL of deionized water into a microtube. |
Comment 1
|

|
| 3) |
Add 2 μL of Endo-H solution to attain a final enzyme concentration of 0.5~10 U /mL. |
Comment 1
|

|
| 5) |
Add 50 μL of 2X SDS-PAGE sample buffer, and then heat at 100°C for 3 min. |
Comment 0
|

|
| 6) |
Load 10 µL of the sample on SDS-PAGE gel and run the electrophoresis. Perform either Coomassie blue staining or silver staining. |
Comment 0
|
|
|
| Figure & Legends |
Figure & Legends 
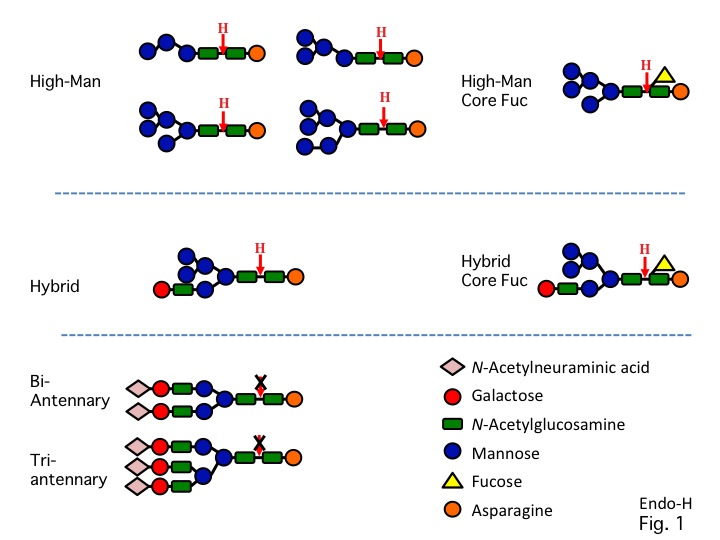
Fig. 1. Specificity of Endo-H. |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2013-11-21 18:29:12 |
- Maley, F., Trimble, R. B., Tarentino, A. L. and Plummer, T. H., Jr. (1989) Characterization of glycoproteins and their associated oligosaccharides through the use of endoglycosidases. Anal. Biochem. 180, 195-204. [PMID : not found]
- Trimble, R. B. and Tarentino, A. L. (1991) Identification of distinct endoglycosidase (endo) activities in Flavobacterium meningosepticum: endo F1, endo F2, and endo F3. Endo F1 and endo H hydrolyze only high mannose and hybrid glycans. J. Biol. Chem. 266, 1646-1651. [PMID : 1899092]
- Tai, T., Yamashita, K., Ogata-Arakawa, M., Koide, N., Muramatsu, T., Iwashita, S., Inoue, Y. and Kobata, A. (1975) Structural studies of two ovalbumin glycopeptides in relation to the endo-beta-N-acetylglucosaminidase specificity. J Biol Chem. 250, 8569-8575. [PMID : 389]
- Tarentino, A. L., Plummer, T. H., Jr. and Maley, F. (1972) A re-evaluation of the oligosaccharide sequence associated with ovalbumin. J Biol Chem. 247, 2629-2631. [PMID : 5019965]
- Tarentino, A. L., Plummer, T. H., Jr. and Maley, F. (1973) A beta-mannosidic linkage in the unit A oligosaccharide of bovine thyroglobulin. J Biol Chem. 248, 5547-5548. [PMID : 4768911]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Fujita, Kiyotaka,
Yamamoto, Kenji,
(2013). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.18,4,2024 .
How to Cite this Work in Website:
Fujita, Kiyotaka,
Yamamoto, Kenji,
(2013).
Endo-β-N-acetylglucosaminidase H digestion (Endo-H).
Retrieved 18,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t71.
html source
Fujita, Kiyotaka,
Yamamoto, Kenji,
(2013).
<b>Endo-β-<em>N</em>-acetylglucosaminidase H digestion (Endo-H)</b>.
Retrieved 4 18,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t71" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t71</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|