This non-RI-based method consists of four steps. First, bacterial sphingomyelinase is used to hydrolyze the sphingomyelin to obtain ceramide and phosphorylcholine. Second, alkaline phosphatase is used to generate choline from the phosphorylcholine. Third, choline oxidase is used to generate hydrogen peroxide (H2O2) from the choline. Finally, H2O2, in the presence of horseradish peroxidase, reacts with dihydroxyphenoxazine (Amplex Red) to generate a highly fluorescent product, resorufin, which is measured by fluorescent microplate reader. |
| Category | Glycolipids and related compounds |
| Protocol Name | Quantitative determination of sphingomyelin II |
Authors
 |
Okino, Nozomu
*
Department of Bioscience and Biotechnology, Graduate School of Bioresource and Bioenvironmental Sciences, Kyushu University
Ito, Makoto
Department of Bioscience and Biotechnology, Graduate School of Bioresource and Bioenvironmental Sciences, Kyushu University
*To whom correspondence should be addressed.
|
| KeyWords |
|
Reagents
 |
| ● |
Reaction buffer: 50 mM Tris-HCl, pH 7.4, containing 5 mM MgCl2 |
| ● |
Standard dilution buffer: 50 mM Tris-HCl, pH 7.4, containing 2% Triton X-100 and 5 mM MgCl2 |
| ● |
12.5 U/mL of Bacillus cereus sphingomyelinase (Sigma-Aldrich, St. Louis, MO, #9396) |
| ● |
400 U/mL of calf intestinal alkaline phosphatase (GE Healthcare, Little Chalfont, UK, E2250Y) |
| ● |
120 U/mL of choline oxidase (Sigma-Aldrich, #26986) |
| ● |
200 U/mL of horseradish peroxidase (Sigma-Aldrich, #77334) |
| ● |
20 mM of Amplex Red (Invitrogen/Life Technologies, Carlsbad, CA, #A12222)
[Note] All stock solutions are prepared in the reaction buffer except Amplex Red, which is in DMSO. |
| ● |
C18-sphingomyelin (Matreya LLC, Pleasant Gap, PA, #1911): Make a 20 mM C18-sphingomyelin stock solution in ethanol and keep at −20°C. |
| ● |
Enzyme cocktail: 12.5 mU of Bacillus cereus sphingomyelinase, 400 mU of alkaline phosphatase, 120 mU of choline oxidase, 200 mU of horseradish peroxidase, and 20 nmol of Amplex Red reagent in 100 μL of reaction buffer |
|
Instruments
 |
| ● |
96-well micro plate (Black plate) |
| ● |
ARVO fluorescence microplate reader (PerkinElmer, Waltham, MA) |
|
| Methods |
|
1. |
Quantitative determination of sphingomyelin [II] |
| 1) |
Homogenize the sample in a 20-fold volume of 0.2% Triton X-100. |
Comment 0
|

|
| 2) |
Centrifuge at 10,000 × g for 5 min. |
Comment 0
|

|
| 4) |
Add 100 μL of enzyme cocktail to each well of a 96-well microtiter plate. |
Comment 0
|

|
| 5) |
Add 5 μL of sample to each well. |
Comment 0
|

|
| 6) |
For controls, add 5 μL of the sample to the enzyme cocktail without sphingomyelinase. |
Comment 0
|

|
| 8) |
Measure the fluorescence of the sample in the microtiter plate with a microplate reader (Set excitation and emission wavelengths at 544 and 590 nm, respectively). |
Comment 0
|
|
|
| Notes | Sphingomyelin content is calculated from the test value after subtracting the control value.
Standard sphingomyelin: 2, 4, 10, 20, 40, 100, 200, 400 pmol/μL (10, 20, 50, 100, 200, 500, 1000, 2000 pmol/5 μL/well) |
| Discussion | With this procedure, 20–1000 pmol/well of sphingomyelin can be determined. |
| Figure & Legends |
Figure & Legends 
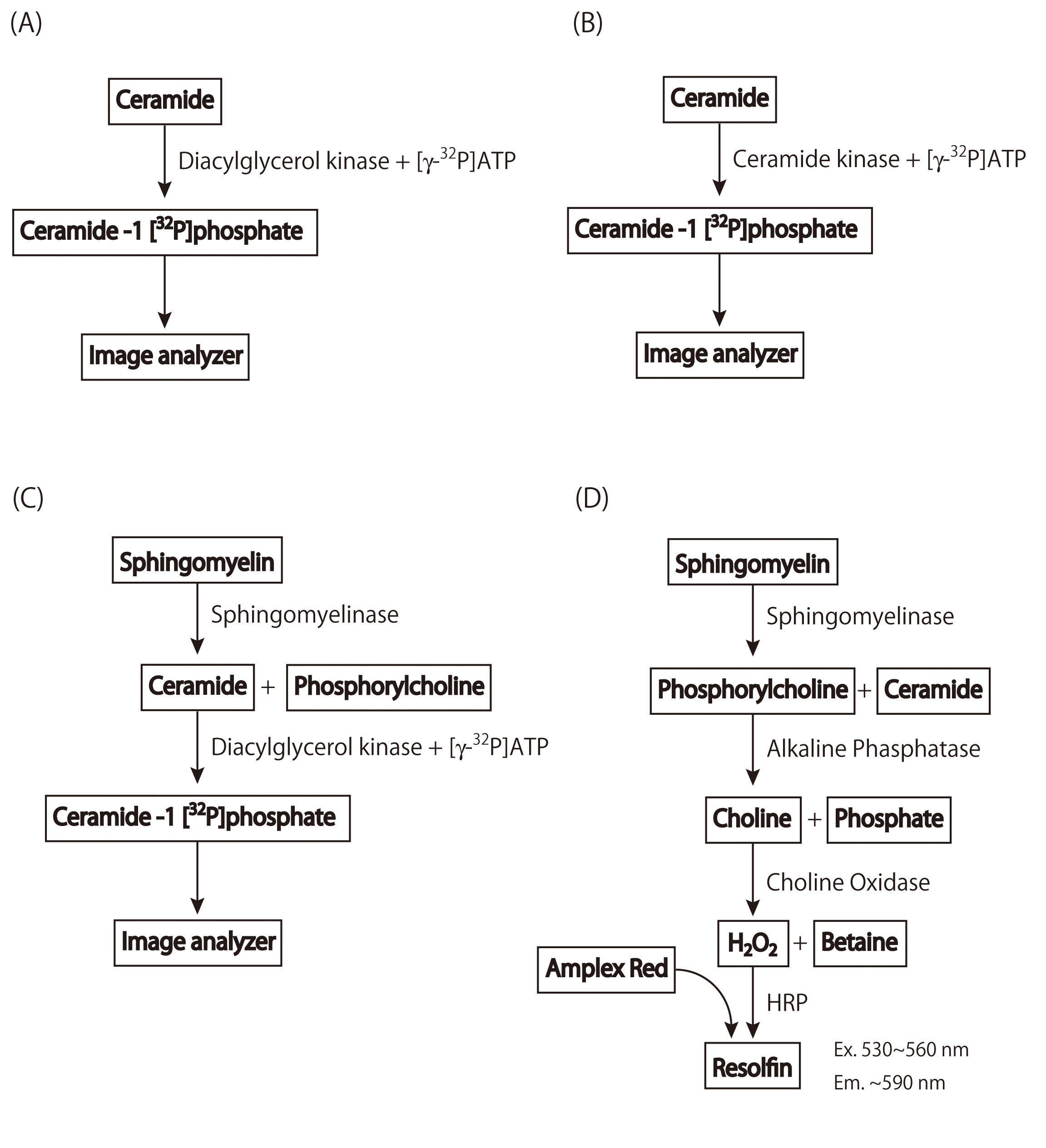
Fig. 1. Scheme for sphingomyelin determination through four enzymatic steps
(A) Quantification of ceramide using diacylglycerol kinase
(B) Quantification of ceramide using ceramide kinase
(C) Quantification of sphingomyelin using sphingomyelinase and diacylglycerol kinase
(D) Quantification of sphingomyelin through four enzymatic steps |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2014-07-31 10:42:46 |
- He, X., Chen, F., McGovern, M.M., and Schuchman, E.H. (2002) A fluorescence-based, high-throughput sphingomyelin assay for the analysis of Niemann-Pick disease and other disorders of sphingomyelin metabolism. Anal. Biochem. 306, 115–123 [PMID : 12069422]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Okino, Nozomu,
Ito, Makoto,
(2014). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.18,4,2024 .
How to Cite this Work in Website:
Okino, Nozomu,
Ito, Makoto,
(2014).
Quantitative determination of sphingomyelin II.
Retrieved 18,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t29.
html source
Okino, Nozomu,
Ito, Makoto,
(2014).
<b>Quantitative determination of sphingomyelin II</b>.
Retrieved 4 18,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t29" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t29</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|