Partial digestion of heparin (Hep)/heparan sulfate (HS) with Hep lyases generates a series of even-numbered oligosaccharides bearing a 4,5-unsaturated uronic acid residue at the non-reducing end. Partial chemical degradation with nitrous acid also generates even-numbered oligosaccharides bearing a 2,5-unhydromannose residue at the reducing end. The resulting oligosaccharides can be size-fractionated by gel filtration chromatography. |
| Category | Glycosaminoglycans |
| Protocol Name | Heparin/heparin sulfate oligosaccharides |
Authors
 |
Yamada, Shuhei
Department of Pathobiochemistry, Faculty of Pharmacy, Meijo University
|
| KeyWords |
|
Reagents
 |
| ● |
Heparinase from Flavobacterium heparinum (Seikagaku Corp., Tokyo, Japan) |
| ● |
Heparitinases from Flavobacterium heparinum (Seikagaku Corp., Tokyo, Japan) |
| ● |
Dimethylmethylene blue dye (Sigma-Aldrich, St. Louis, MO) |
| ● |
Ba(NO2)2 (Nacalai Tesque Inc., Kyoto, Japan) |
|
Instruments
 |
| ● |
Bio-Gel P-10 resin (Bio-Rad Laboratories, Hercules, CA) |
| ● |
Sephadex G-25 fine resin (GE Healthcare, Little Chalfont, UK) |
| ● |
SuperdexTM Peptide 10/300 GL column (GE Healthcare) |
| ● |
Speed Vac Concentrator (Thermo Fisher Scientific Inc., Waltham, MA) |
| ● |
HPLC system (LC-20 Series: Shimadzu Corporation, Kyoto, Japan) |
|
| Methods |
|
1. |
|
| 1) |
Incubate Hep/HS (20 mg) with Hep lyase (heparinase, heparitinase, or a mixture of heparinase and heparitinase) (0.2 IU) in a total volume of 2 mL of 20 mM acetate-Na buffer, pH 7.0, containing 2 mM Ca(CH3COO)2 at 37˚C. |
Comment 1
|

|
| 2) |
Withdraw aliquots (5 μL) at 0.5- to 1- h intervals to monitor the reaction. |
Comment 0
|

|
| 3) |
Mix the aliquot with 0.01 M HCl (495 μL) and measure the UV-absorbance at 232 nm. |
Comment 1
|

|
| 4) |
Terminate the reaction by boiling at 100˚C for 2 min, when the absorbance reaches the target value. |
Comment 1
|

|
| 5) |
Fractionate the digest by gel filtration chromatography on a Bio-Gel P-10 or SuperdexTM Peptide column. |
Comment 1
|

|
| 6) |
Desalt the subfractions by gel filtration chromatography on a Sephadex G-25 fine column using distilled water as the eluent. |
Comment 1
|

|
| 7) |
Lyophilize the flow-through fraction by Speed Vac. |
Comment 0
|
|
|
|
2. |
Chemical degradation of Hep/HS. (To chemically depolymerize Hep/HS chains, nitrous acid treatment is often used.) |
| 1) |
Lyophilize Hep/HS (50 μg) in a glass tube by Speed Vac. |
Comment 0
|

|
| 2) |
Add 40 μL of 0.5 M HNO2 (pH 1.5) reagent and incubate at room temperature for 30 min. |
Comment 0
|

|
| 3) |
Neutralize the incubation mixture with 1 M NaOH. |
Comment 1
|

|
| 4) |
Add 50 μL of 1 M NaBH4/0.05 M NaOH and incubate at room temperature for 2 h. |
Comment 1
|

|
| 5) |
Acidify the incubation mixture with glacial acetic acid to terminate the reaction. |
Comment 1
|

|
| 7) |
Fractionate oligosaccharides by gel filtration chromatography on a SuperdexTM Peptide column. |
Comment 1
|
|
|
| Figure & Legends |
Figure & Legends

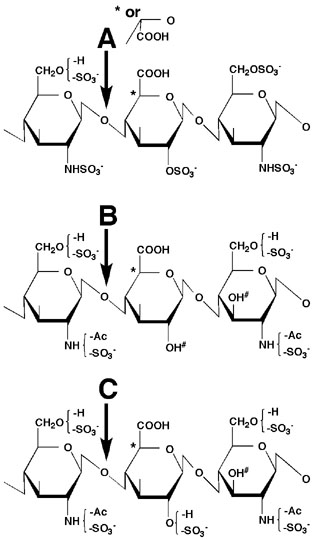
Fig. 1. Substrate specificity of Hep lyases
The arrows indicate the glycosidic linkages cleaved by heparinase (A), heparitinase I (B), and heparitinase II (C). The hexuronate residues (marked by asterisks) are either D-glucuronic acid (GlcA) or L-iduronic acid (IdoA). C-2 NH2 of the D-glucosamine (GlcN) and C-2 OH of the hexuronate residue should be sulfated for sensitivity to heparinase (A). C-3 OH of GlcN adjacent to the disaccharide cleavage site (marked by #) has to be free for sensitivity to heparitinases I and II (B, C). Heparitinase I recognizes nonsulfated hexuronate (C-2 OH should be free as marked by #) and prefers GlcA to IdoA as a substrate (B).

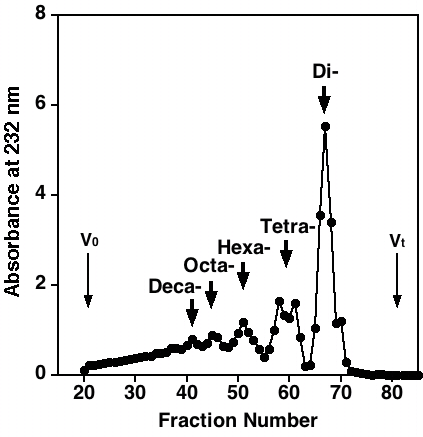
Fig. 2. Fractionation of the heparitinase digest of bovine kidney HS
Bovine kidney HS (33 mg) was digested by heparitiase until the reaction reached 60% completion. The digest was gel-filtrated on a column (1.6 cm i.d. x 95 cm) of Bio-Gel P-10 using 1.0 M NaCl/10% ethanol as the eluent. Two-milliliter fractions were collected and aliquots were monitored by measuring absorbance at 232 nm. The sizes of oligosaccharides in the resolved peaks are indicated.
This figure was originally published in Glycobiology. 4(4):535-44. 1994 "Structural studies on the oligosaccharides isolated from bovine kidney heparan sulphate and characterization of bacterial heparitinases used as substrates" Sugahara K. Yamada S. et al. Oxford University Press.

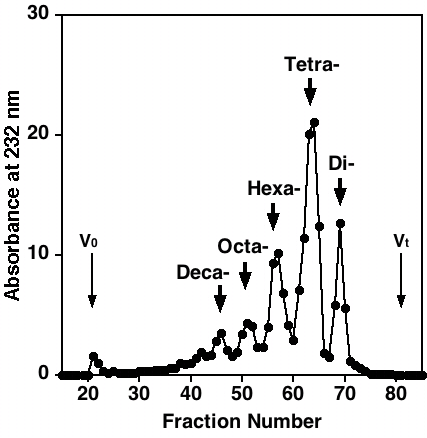
Fig. 3. Gel-filtration of the heparinase digest on Bio-Gel P-10
Hep (300 mg) was digested with heparinase. The digest was subjected to gel-filtration on a column (1.5 cm i.d. x 46 cm) of Sephadex G-25 fine using 0.25 M NH4HCO3/7% 1-propanol as the eluent, and separated into oligosaccharides and disaccharides (data not shown). The oligosaccharide fraction was further fractionated on a column (1.6 cm i.d. x 95 cm) of Bio-Gel P-10 using 1.0 M NaCl/10% ethanol as the eluent. Fractions (2 mL) were collected and monitored by measuring absorbance at 232 nm. The sizes of oligosaccharides in the resolved peaks are indicated.
This figure was originally published in J Biol Chem. Yamada S. et al. "Isolation of the porcine heparin tetrasaccharides with glucuronate 2-O-sulfate. Heparinase cleaves glucuronate 2-O-sulfate-containing disaccharides in highly sulfated blocks in heparin" 1995, 270(15):8696-705. © the American Society for Biochemistry and Molecular Biology. |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2013-12-27 15:45:37 |
- Sugahara, K., Tohno-oka, R., Yamada, S., Khoo, K-H., Morris, H.R., and Dell, A. (1994) Structural studies on the oligosaccharides isolated from bovine kidney heparan sulphate and characterization of bacterial heparitinases used as substrates. Glycobiology 4, 535-544. [PMID : 7827415]
- Yamada, S., Murakami, T., Tsuda, H., Yoshida, K., and Sugahara, K. (1995) Isolation of the porcine heparin tetrasaccharides with glucuronate 2-O-sulfate: heparinase cleaves glucuronate 2-O-sulfate-containing disaccharides in highly sulfated blocks in heparin. J Biol Chem 270, 8696-8705. [PMID : 7721774]
- Kreuger, J., Prydz, K., Pettersson, R.F., Lindahl, U., and Salmivirta, M. (1994) Characterization of fibroblast growth factor 1 binding heparan sulfate domain. Glycobiology 9, 723-729. [PMID : 10362842]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Yamada, Shuhei,
(2013). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.18,4,2024 .
How to Cite this Work in Website:
Yamada, Shuhei,
(2013).
Heparin/heparin sulfate oligosaccharides.
Retrieved 18,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t18.
html source
Yamada, Shuhei,
(2013).
<b>Heparin/heparin sulfate oligosaccharides</b>.
Retrieved 4 18,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t18" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t18</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|