Oligosaccharide linked to glycoprotein or glycopeptide plays important roles in several biological events. However, oligosaccharide exhibits heterogeneity and this has been a hindrance in order to investigate what oligosaccharide is essential for the several biological events. Therefore, it is essential to synthesize glycoproteins and glycopeptides having homogeneous oligosaccharides. This protocol will show synthetic method of glycoprotein by native chemical ligation. |
| Category | Large scale preparation of glycans, glycoproteins & glycolipids |
| Protocol Name | Synthesis of the glycoprotein having N-linked complex type oligosaccharide |
Authors
 |
Okamoto, Ryo
Department of Chemistry, Osaka University
Kajihara, Yasuhiro
*
Department of Chemistry, Osaka University
*To whom correspondence should be addressed.
|
| KeyWords |
|
Reagents
 |
| ● |
4-(4-Hydroxymethyl-3-methoxyphenoxy)-butyric acid (HMPB)-PEGA Resin |
| ● |
9-Fluorenylmethyl-carbonate (Fmoc)-amino acid derivatives |
| ● |
1-Mesitylenesulfonyl-3-nitro-1,2,4-triazole (MSNT) |
| ● |
|
| ● |
|
| ● |
3-(Diethoxy-phosphoryloxy)-1,2,3-benzotriazin-4(3H)-one (DEPBT) |
| ● |
Diisopropylcarbodiimide (DIC) |
| ● |
1- Hydroxybenzotriazole (HOBt) |
| ● |
|
| ● |
Trifluoroacetic acid (TFA) |
| ● |
|
| ● |
N,N’-Diisopropylethylamine (DIEA) |
|
Instruments
 |
| ● |
HPLC system and a reverse phase column (Cadenza column, Imtakt Corp., Kyoto, Japan, 3 mm, 75 × 4.6 mm, at a flow rate of 1.0 mL min-1). |
|
| Methods |
|
1. |
Synthesis of sialylglycopeptide-benzylthioester 4, QPVGIN(Bn-CHO)TSTT-SBn (Fig. 1)1) |
| 1) |
Couple Fmoc-Thr-(OtBu)-OH (3 equiv.) to the HMPB-PEGA resin using MSNT (3.0 equiv.) and N-methylimidazole (2.75 equiv.) in CH2Cl2 (concentration of Fmoc-Thr(OtBu)-OH was adjusted to 250 mM). |
Comment 0
|

|
| 2) |
Add 20% piperidine in DMF and react for 20 min to deprotect the Fmoc group. |
Comment 0
|

|
| 3) |
Couple the amino acid by use of Fmoc-amino acid derivatives (5.0 equiv.) according to the target amino acid sequence, diisopropylcarbodiimide (DIC, 5.0 equiv.) and 1-hydroxybenzotriazole (HOBt, 5.0 equiv.) in DMF (concentration of Fmoc-amino acid was adjusted to 0.4 M) and react for 1.0 h. Deprotect the Fmoc group by the treatment with 20% piperidine in DMF for 20 min after each Fmoc-amino acid coupling. |
Comment 0
|

|
| 4) |
Add the Fmoc-Asn(oligosaccharide)-OH 1 (2.0 equiv.) to the peptide-resin with DEPBT (3.0 equiv.) and DIPEA (2.0 equiv.) in DMF (concentration of 1 was adjusted to 30 mM), followed by deprotection of the Fmoc group by the treatment with 20% piperidine in DMF for 20 min. |
Comment 0
|

|
| 5) |
Repeat the same manner as step 3 until the last amino acid residue but adjust the concentration of the Fmoc-amino acid to 40 mM. |
Comment 0
|

|
| 6) |
Add the solution of acetic acid : trifluoroethanol (1 : 1, e.g. 2.0 mL for 2 micro mol scale glycopeptide synthesis) and repeat this twice in order to obtain side chain protected glycopeptide 2. |
Comment 0
|

|
| 7) |
Concentrate the solution containing 2 in vacuo. |
Comment 0
|

|
| 8) |
Dissolve the residue in DMF and concentrate in vacuo for three times. |
Comment 0
|

|
| 9) |
Dissolve the residue containing the glycopeptide 2 in DMF (the concentration was adjusted to 5 mM). |
Comment 0
|

|
| 10) |
Add 4 Å molecular sieves (10 mg for 2 micro mol scale glycopeptide synthesis) and benzylthiol (30 equiv.) to the DMF solution and incubate for 1 h at −20˚C. |
Comment 0
|

|
| 11) |
Add PyBOP (5 equiv.) and DIPEA (5 equiv.) to this mixture and stirr the mixture at −20˚C for 4 h. |
Comment 0
|

|
| 12) |
Filter the solution and add ethyl ether (e.g. 6.0 mL for 2 micro mol scale glycopeptide synthesis) to give a precipitation of glycopeptide-alpha-thioester 3. Collect the precipitate by a centrifugation. |
Comment 0
|

|
| 13) |
Add a solution containing 95% TFA, 2.5% triisopropyl silane (TIPS), and 2.5% H2O to the precipitate to remove side-chain protecting groups. After 2 h, concentrate the solution in vacuo. |
Comment 0
|

|
| 14) |
Purifiy the residue by reverse phase-HPLC to obtain sialylglycopeptide-alpha-benzylthioester 4 (3.0 mg, 43%).
1H-NMR (400 MHz, 295K in D2O, HOD = δ 4.81) δ 7.54-7.35 (m, 15H, Ph,), 5.38 (d, 2H, J = 11.8 Hz, PhCH2), 5.28 (d, 2H, J = 11.8 Hz, PhCH2), 5.10 (s, 1H, Man4-H-1), 5.01 (d, 1H, J = 9.6 Hz, GlcNAc1-H-1), 4.92 (s, 1H, Man4’-H-1), 4.77 (s, 1H, Man3-H-1), 4.70 (dd, 1H, Pyr-αH), 4.25 (bs, 1H, Man3-H-2), 2.91 (bdd, 1H, Asn-βCH2), 2.75 (bdd, 1H, Asn-βCH2), 2.65 (bdd, 2H, NeuAc7, 7’-H3eq), 1.84 (m, 3H, Ile-β CH2, NeuAc7, 7’-H-3ax), 1.23, 1.15, 1.13 (each d, each 3H, Thr-γCH3), 1.97, 1.93 (each d, each 3H, Val-γCH3), 1.86 (d, 3H, Ile-γCH3), 1.82 (dd, 3H, Ile-γCH3) ; ESI-MS m/z calcd for [M+H]+ 3490.4, found 3490.2 (deconvoluted). |
Comment 0
|
|
|
|
2. |
Synthesis of glycosylated MCP-3 using sequential Native Chemical Ligation (Fig. 2)1) |
| 1) |
Dissolve the peptide segment-2 (5, 2 mM) and segment-3 (6, 2 mM) in 0.1 M phosphate buffer (pH 7.5) solution containing 6 M Gn-HCl, 1% thiophenol (v/v) and 1% benzylmercaptan (v/v). |
Comment 0
|

|
| 2) |
Adjust the pH of the reaction solution to 4.0 by using 0.2 M methoxyamine-HCl to convert the N-terminal Thz-group to Cys residue. |
Comment 0
|

|
| 3) |
Purify the reaction mixture by reverse phase-HPLC to obtain polypeptide segment-4 (7) (ESI; m/z calcd for [M+H]+ 7854.1, found 7855.1 (deconvoluted)). |
Comment 0
|

|
| 4) |
Dissolve the peptide segment-4 (7, 2 mM) and segment-1 (sialylglycopeptide-alpha-benzylthioester 4, 2 mM) in 0.1 M phosphate buffer (pH 7.6) solution containing 6 M Gn-HCl, 1% thiophenol (v/v) and 1% benzylmercaptan (v/v) and incubate for 17 h at 37˚C. |
Comment 0
|

|
| 5) |
Add the sialylglycopeptide-alpha-benzylthioester 4 (ca. 0.5–0.6 equiv. to the used segment-4) to the reaction mixture, and incubate this mixture for 17 h at 37˚C. |
Comment 0
|

|
| 6) |
Dilute the mixture by adding five times volume of 0.1M Tris/HCl buffer (pH 8.0) and then bubble the air for 1min to perform folding reaction. Incubate the reaction mixture for 24 h at room temperature. |
Comment 0
|

|
| 7) |
Purify the mixture by reverse phase-HPLC to afford the desired product (ESI; m/z calcd for [M+7H]7+ 1617.4, [M+8H]8+ 1415.3, [M+9H]9+ 1258.2, [M+10H]10+ 1132.4, [M+11H]11+ 1029.6, [M+12H]12+ 943.9, found 1617.5, 1415.5, 1258.4, 1132.7, 1029.9, 944.2). |
Comment 0
|

|
| 8) |
Add the 50 mM NaOH and leave for 10 min to remove the benzyl ester groups. |
Comment 0
|

|
| 9) |
Purify the mixture by reverse phase-HPLC to afford glycosylated MCP-3 9 ESI; m/z calcd for [M+7H]7+ 1591.6, [M+8H]8+ 1392.8, [M+9H]9+ 1238.1, [M+10H]10+ 1114.4, [M+11H]11+ 1013.2, [M+12H]12+ 928.9, found 1591.8, 1393.1, 1238.5, 1114.7, 1013.5, 929.1). |
Comment 0
|
|
|
| Notes | For the glycopeptide synthesis, the concentrations of Fmoc-amino acids following the Fmoc-Asn(oligosaccharide)-OH were arranged to be 40 mM in DMF in order to avoid unexpected esterification toward hydroxyl groups on the oligosaccharide. |
| Figure & Legends |
Figure & Legends 
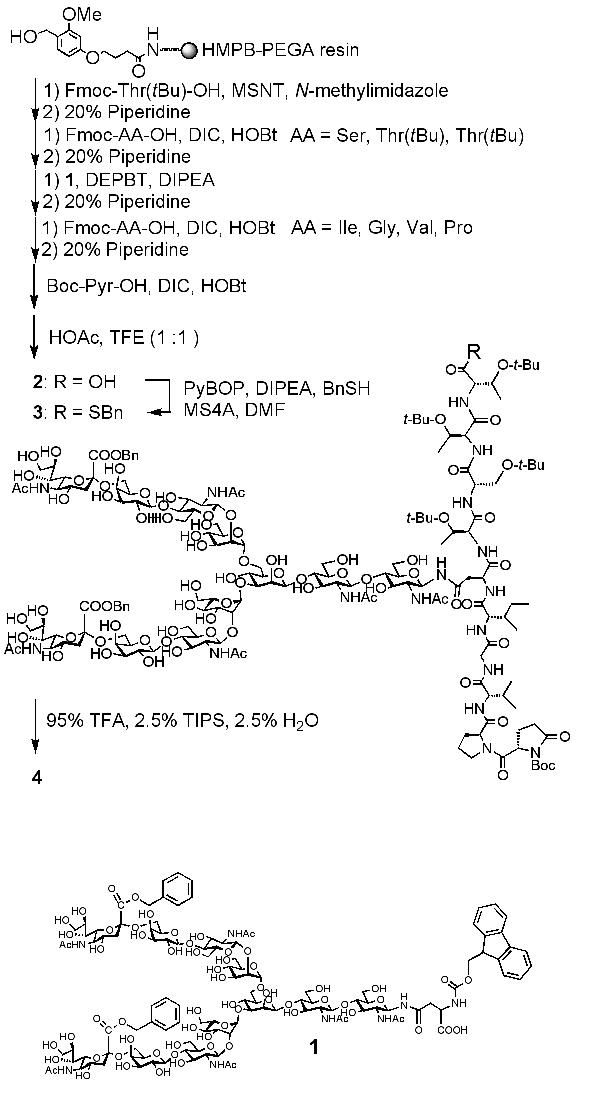
Fig. 1

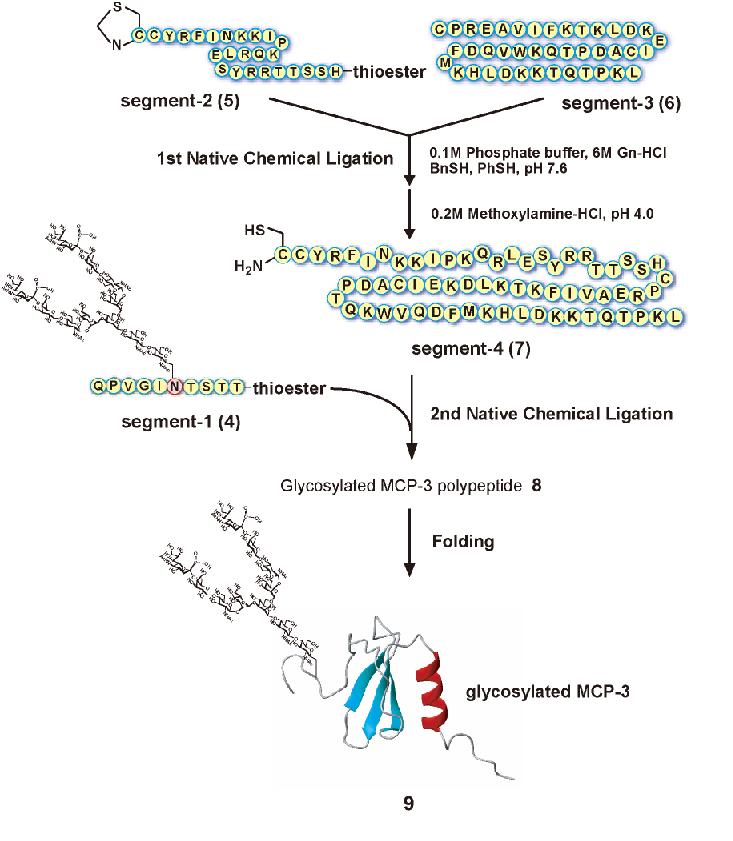
Fig. 2 |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2015-01-20 13:19:03 |
- Yamamoto, N., Tanabe, Y., Okamoto, R., Dawson, P.E., Kajihara Y. (2008) Chemical synthesis of a glycoprotein having an intact human complex-type sialyloligosaccharide under the Boc and Fmoc synthetic strategies. J. Am. Chem. Soc. 130, 501–510 [PMID : 18085777]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Okamoto, Ryo,
Kajihara, Yasuhiro,
(2015). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.25,4,2024 .
How to Cite this Work in Website:
Okamoto, Ryo,
Kajihara, Yasuhiro,
(2015).
Synthesis of the glycoprotein having N-linked complex type oligosaccharide.
Retrieved 25,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t116.
html source
Okamoto, Ryo,
Kajihara, Yasuhiro,
(2015).
<b>Synthesis of the glycoprotein having <em>N</em>-linked complex type oligosaccharide</b>.
Retrieved 4 25,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t116" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t116</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|