PNGase (peptide:N-glycanase or peptide-N4-(N-acetyl-β-D-glucosaminyl)asparagine amidase (EC 3.5.1.52)) cleaves the amide bond between proximal GlcNAc and the linkage Asn residue of N-linked glycoproteins. PNGases from plant or bacterial origin have been widely used as powerful tool reagents for analyzing the structures and functions of N-glycans on glycoproteins. The cytoplasmic PNGase was found to be quite distinct in terms of enzymatic properties from the reagent PNGases [Suzuki, et al. J. Biol. Chem. 1994; Suzuki, et al. J. Biol. Chem. 1995].
It has been established that eukaryotic cells have a so-called “ER-associated degradation (ERAD)” system for elimination of newly synthesized misfolded/unassembled proteins. So far a number of ERAD substrates are known to be deglycosylated by the cytoplasmic PNGase during their degradation [Tanabe, et al. Methods Enzymol. 2006 and references therein].
Deglycosylation of misfolded glycoproteins in vivo can be routinely monitored by SDS-PAGE/Western blotting using substrates of your choices [ex. Tanabe, et al. Methods Enzymol. 2006]. |
| Category | Biosynthesis & Metabolism |
| Protocol Name | Assay for cytoplasmic PNGase |
Authors
 |
Suzuki, Tadashi
Glycometabolome Team, RIKEN Global Research Cluster
|
| KeyWords |
|
Reagents
 |
| ● |
RNase B (Sigma-Aldrich, St. Louis, MO, R5870) |
| ● |
|
| ● |
|
| ● |
|
| ● |
1 M Tris-HCl buffer (pH 7.5/8.0) |
| ● |
|
| ● |
0.5 M EDTA (pH adjusted with 1 N NaOH to 8.0) |
| ● |
AESMF (Pefabloc SC: Roche Applied Science, Penzberg, Germany) |
| ● |
CompleteTM protease inhibitor cocktails (Roche Applied Science) |
| ● |
Anti-RNase antibody (Rockland Immunochemicals, Gilbertsville, PA, Code No. 200-4188) |
| ● |
Standard reagents for SDS-PAGE/Western blotting |
|
Instruments
 |
| ● |
Equipments for mammalian cell culture |
| ● |
|
| ● |
|
| ● |
|
| ● |
Devices for SDS-PAGE/Western blotting |
| ● |
|
|
| Methods |
|
1. |
Isolation of S-alkylated RNase B [Suzuki, Methods 2005] |
| 1) |
Dissolve 20 mg of RNase B in 2 mL of 8 M urea/10 mM DTT/0.1 M Tris-HCl buffer (pH 8.0). Incubate at 37°C for 30 min. |
Comment 0
|

|
| 2) |
Add 174 μL of 1M iodoacetamide in 8 M urea/10 mM DTT/0.1 M Tris-HCl buffer (pH 8.0). Final concentration of iodoacetamide will be 80 mM. |
Comment 0
|

|
| 3) |
Incubate further at 37°C for 30 min. Avoid light during incubation. |
Comment 0
|

|
| 4) |
Pass the sample through a PD-10 column (GE Healthcare, Little Chalfont, UK) to exchange the buffer with 50 mM NaCl/20 mM Tris/HCl buffer (pH 7.5). |
Comment 0
|

|
| 5) |
Adjust the protein concentration to ~5 mg/mL with the elution buffer. The substrates prepared can be aliquoted (~50 μL), stored at −20°C or lower until use. |
Comment 0
|
|
|
|
2. |
Preparation of Cytoplasmic fraction for enzyme source |
| 1) |
Culture cells of your interest. Collect cells. |
Comment 0
|

|
| 2) |
Perform all the subsequent procedures either on ice or at 4°C. Suspend cells at a density of 5 × 107 cells/ml in 10 mM Tris-HCl buffer (pH 7.5)/1 mM EDTA/250 mM sucrose/1 mM DTT with various protease inhibitors (1 × completeTM protease inhibitor cocktail/1 mM AEBSF (Pefabloc SC)). |
Comment 0
|

|
| 3) |
Homogenize cells using Potter-Elvehjem homogenizer or equivalent. |
Comment 0
|

|
| 4) |
Clear the solution first with regular centrifuge at 14,000 rpm for 10 min. |
Comment 0
|

|
| 6) |
Centrifuge the sup further using ultracentrifuge at 100,000 × g for 1 h. |
Comment 0
|

|
| 7) |
Soluble (cytosol) fraction thus obtained can be used for PNGase assay. For later use, the sample can be aliquoted, stored at −80°C. Avoid repeated freeze-thawing. |
Comment 0
|
|
|
|
3. |
Assay for PNGase activity using S-alkylated RNase B as a substrate |
| 1) |
Take 9 μL of the cytosol fraction, mix with 1 μL of S-alkylated RNase B solution prepared as described above. |
Comment 0
|

|
| 2) |
Incubate at 30°C. When cytosol was used as an enzyme source, incubation time has to be determined empirically; as an initial attempt overnight incubation is recommended. |
Comment 0
|

|
| 3) |
Run the sample on 15% SDS-PAGE gel (mix with 2X sample buffer and run 10 μL (ca. 3 μg for RNase B) for each sample). |
Comment 0
|

|
| 4) |
Carry out Western blot analysis using rabbit anti-RNase antibody. |
Comment 0
|

|
| 5) |
Visualize the gel using fluorescent gel imager. |
Comment 0
|
|
|
| Notes | If significant amount of nonglycosylated RNase (RNase A) is contaminated in the commercial preparation of RNase B, purification of RNase B can be achieved using Concanavalin A-Sepharose column, as described previously [Suzuki, T (2005) Methods].
For PNGase-reaction, incubation at higher temperature (37°C) often caused rapid inactivation of the enzyme and therefore not recommended. |
| Figure & Legends |
Figure & Legends 
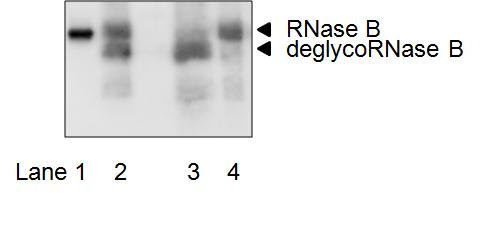
Fig. 1.
For enzyme source, CHO-K1 cells were used and incubation was carried out at 30°C for 15 h. Similar results can be obtained for other mammalian-derived culture cells. SDS-PAGE gel was visualized using LAS-3000 mini (Fujifilm, Tokyo, Japan). Antibody dilution: primary antibody; 1/1,000 (rabbit anti-RNase antibody, Rockland Immunochemicals); secondary antibody; 1/2,000 (donkey horseradish peroxidase-linked anti-rabbit IgG antibody, GE Healthcare).
Standard deglycosylated sample (lane 3) was obtained by digesting 25 μg of S-alkylated RNase B with 2.5U PNGase F (Roche Applied Science) in 25 μL of 80 mM sodium phosphate buffer (pH 7.0) at 37°C for 15 h.
Lane 1, Molecular Marker (20K; Magic MarkTM XP Western Protein Standard, Invitrogen/Life Technologies, Carlsbad, CA);
Lane 2, Reaction mixture from the cytosol of CHO-K1 cells
Lane 3, Control deglycosylated sample
Lane 4, Control Substrate (S-alkylated RNase B).
RNase B: S-alkylated RNase B (substrate);
deglycoRNase B (de-N-glycosylated S-alkylated RNase B (PNGase reaction product)). |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2014-12-16 09:32:24 |
- Suzuki, T., Seko, A., Kitajima, K., Inoue, Y., and Inoue, S. (1994) Purification and enzymatic properties of peptide: N-glycanase from C3H mouse-derived L-929 fibroblast cells. Possible widespread occurrence of post-translational remodification of proteins by N-deglycosylation. J. Biol. Chem. 269, 17611–17618 [PMID : 8021270]
- Suzuki, T., Kitajima, K., Inoue, Y., and Inoue, S. (1995) Carbohydrate-binding property of peptide: N-glycanase from mouse fibroblast L-929 cells as evaluated by inhibition and binding experiments using various oligosaccharides. J. Biol. Chem. 270, 15181–16186 [PMID : 7797502]
- Suzuki, T. (2005) A simple, sensitive in vitro assay for cytoplasmic deglycosylation by peptide: N-glycanase. Methods 35, 360–365, 2005 [PMID : 15804608]
- Tanabe, K. Lennarz, W.J. and Suzuki, T. (2006) A cytoplasmic peptide: N-glycanase. Methods Enzymol. 415, 46–55 [PMID : 17116467]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Suzuki, Tadashi,
(2014). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.20,4,2024 .
How to Cite this Work in Website:
Suzuki, Tadashi,
(2014).
Assay for cytoplasmic PNGase.
Retrieved 20,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t108.
html source
Suzuki, Tadashi,
(2014).
<b>Assay for cytoplasmic PNGase</b>.
Retrieved 4 20,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t108" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t108</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|