Galectin-9, which consists of two carbohydrate recognition domains joined by a linker peptide, belongs to the tandem-repeat-type subclass of the galectin family. Alternative splicing leads to the formation of at least three distinct splice variants (isoforms) of galectin-9 with tandem-repeat-type structures. The isoforms share identical carbohydrate recognition domains, differing only in the linker peptide region. Galectin-9, like other galectins, does not have a signal sequence and is mainly localized in the cytoplasm. However, it is secreted into the extracellular space through unconventional mechanisms and exerts its biological functions by binding to glycoconjugates on the target cell surface.
Human galectin-9 was first identified as a novel tumor antigen of unknown function in patients with Hodgkin’s disease. Subsequently, an independent study identified galectin-9 (ecalectin) as a T cell-derived eosinophil chemoattractant. This study was followed by many others demonstrating the immunomodulatory function of the molecule. Currently, galectin-9 is considered to be a novel type of regulator of the immune response and homeostasis. |
| Category | Sugar binding proteins |
| Protocol Name | Expression and purification of recombinant human galectin-9 |
Authors
 |
Nishi, Nozomu
*
Division of Research Instrument and Equipment, Life Science Research Center, Kagawa University
Nakamura, Takanori
Department of Endocrinology, Faculty of Medicine, Kagawa University
*To whom correspondence should be addressed.
|
| KeyWords |
|
Reagents
 |
| ● |
E. coli (BL21(DE3)) cells carrying an expression plasmid for human galectin-9* (glycerol stock)
* The coding region of human galectin-9 cDNA being inserted into the Nde I-Bam HI sites of the pET-11a vector. |
| ● |
LB-broth containing 100 μg/mL ampicillin
- 2xYT medium containing 100 μg/mL ampicillin
- 0.1 M isopropyl-β-thiogalactopyranoside (IPTG)
|
| ● |
10 mM Tris-HCl (pH 7.5), 0.5 M NaCl, 1 mM phenylmethylsulfonyl fluoride (PMSF) (E. coli suspension buffer) |
| ● |
|
| ● |
20 mM Tris-HCl (pH 7.5), 0.15 M NaCl (TBS), 0.03%
3-[(3-cholamidopropyl) dimethylammonio]-1-propanesulfonate (CHAPS) (lactose-agarose column washing buffer) |
| ● |
TBS, 0.2 M lactose (lactose-agarose column elution buffer) |
| ● |
Phosphate-buffered saline (PBS) (dialysis buffer) |
| ● |
Lactose-agarose slurry (50%[v/v] in TBS) |
|
Instruments
 |
|
| Methods |
|
1. |
Expression and purification of recombinant human galectin-9 |
| 1) |
Inoculate 5 mL of LB-broth containing 100 μg/mL ampicillin with a loop full of frozen BL21(DE3) cells carrying the expression plasmid. |
Comment 0
|

|
| 2) |
Incubate the inoculum at 37°C overnight with shaking (200 rpm) in the shaking incubator. |
Comment 0
|

|
| 3) |
Dilute 4 mL of the overnight culture with 200 mL* of 2xYT medium containing 100 μg/mL ampicillin. |
Comment 1
|

|
| 4) |
Incubate the inoculum at 37°C for 1.5–2.5 h with shaking until the absorbance at 600 nm (A600 nm) reaches 0.6–0.7. |
Comment 0
|

|
| 5) |
Add 0.2 mL of 0.1 M IPTG solution to the culture (final concentration, 0.1 mM). |
Comment 0
|

|
| 6) |
Incubate the culture at 20°C* overnight with shaking. |
Comment 1
|

|
| 7) |
Transfer the overnight culture to centrifuge tubes and then spin down the cells at 6,000 × g for 10 min. |
Comment 0
|

|
| 8) |
Resuspend the pellet in 36 mL (18% of the original culture volume) of 10 mM Tris-HCl (pH 7.5), 0.5 M NaCl, 1 mM PMSF. |
Comment 1
|

|
| 9) |
Transfer the cell suspension to a cooling cell and then lyse the E. coli cells using the sonicator. |
Comment 1
|

|
| 10) |
Add 4 mL of 10% (w/v) Triton X-100 to the cell lysate (final concentration, 1%). |
Comment 0
|

|
| 11) |
Mix the lysate for 30 min with a magnetic stirrer. |
Comment 0
|

|
| 12) |
Transfer the lysate to centrifuge tubes and then spin down the cell debris at 18,000 × g for 30 min. |
Comment 0
|

|
| 13) |
Transfer the supernatant to a 100 mL bottle. |
Comment 0
|

|
| 14) |
Add 1 mL of the lactose-agarose slurry (50%[v/v] in TBS) to the supernatant (0.5 mL of lactose-agarose gel per 200 mL of the original culture volume). |
Comment 0
|

|
| 15) |
Mix the lactose-agarose gel suspension by rotating the bottle for 1 h on the tube rotator. |
Comment 1
|

|
| 16) |
Transfer the gel suspension to a 50 mL conical tube and then spin down the gel at 1,500 × g for 5 min. |
Comment 0
|

|
| 17) |
Discard the supernatant and then suspend the gel pellet in an appropriate volume of TBS, 0.03% CHAPS. |
Comment 0
|

|
| 19) |
Wash the gel with at least 10 gel-bed volumes of TBS, 0.03% CHAPS. |
Comment 0
|

|
| 20) |
Elute the recombinant protein from the gel with TBS, 0.2 M lactose. |
Comment 1
|

|
| 21) |
Dialyze the eluate against PBS. |
Comment 0
|

|
| 22) |
Transfer the dialyzed solution to a centrifuge tube and then spin down the insoluble material at 25,000 × g for 20 min. |
Comment 0
|

|
| 23) |
Sterilize the supernatant with a disposable filter (0.2 μm). |
Comment 1
|

|
| 24) |
Store the sterilized solution at 4°C. |
Comment 1
|
|
|
| Notes | Galectin-9 is a "difficult-to-express protein" among the members of the galectin family. We have evaluated a variety of expression systems including E. coli cells, yeast (S. cerevisiae and P. pastoris) cells, moss cells (Physcomitrella patens), baculovirus-infected insect cells, and some mammalian cells as to their ability to produce recombinant galectin-9. As a result, we found that the E. coli expression system (1st choice: tag-free expression using a pET system; 2nd choice: GST fusion system) is best suited for the production of the protein, although the yield is low compared to those of other members of the family. The low yield is partly due to the low solubility of galectin-9: the maximum concentration of galectin-9 attainable in PBS (and other solvents compatible with biological experiments) is typically lower than 0.4 mg/mL.
There are at least three isoforms of galectin-9 with tandem-repeat-type structures, G9S, G9M and G9L, in order of the length of the linker peptide, G9S being the isoform with the shortest one. The isoforms share identical carbohydrate recognition domains, differing only in the linker peptide region. The isoforms are indistinguishable as to biological activity so far examined. The yields of the recombinant proteins are inversely related to the length of the linker peptide. In addition, G9M and G9L are highly susceptible to proteolysis due to the presence of the relatively long linker peptide. Accordingly, G9S is the isoform of choice for expression in E. coli cells, although it is a minor one (G9M is the predominant isoform in most cell and tissue types). When using G9M/L, the addition of protease inhibitors (preferably a protease inhibitor cocktail) to the final preparation is indispensable for storage for longer than a few days. Galectin-8, another member of the tandem-repeat-type subclass of the galectin family, can be expressed and purified by essentially the same procedure as described above. The average yield of galectin-8 is about 20 times higher than that of G9S. |
| Figure & Legends |
Figure & Legends 
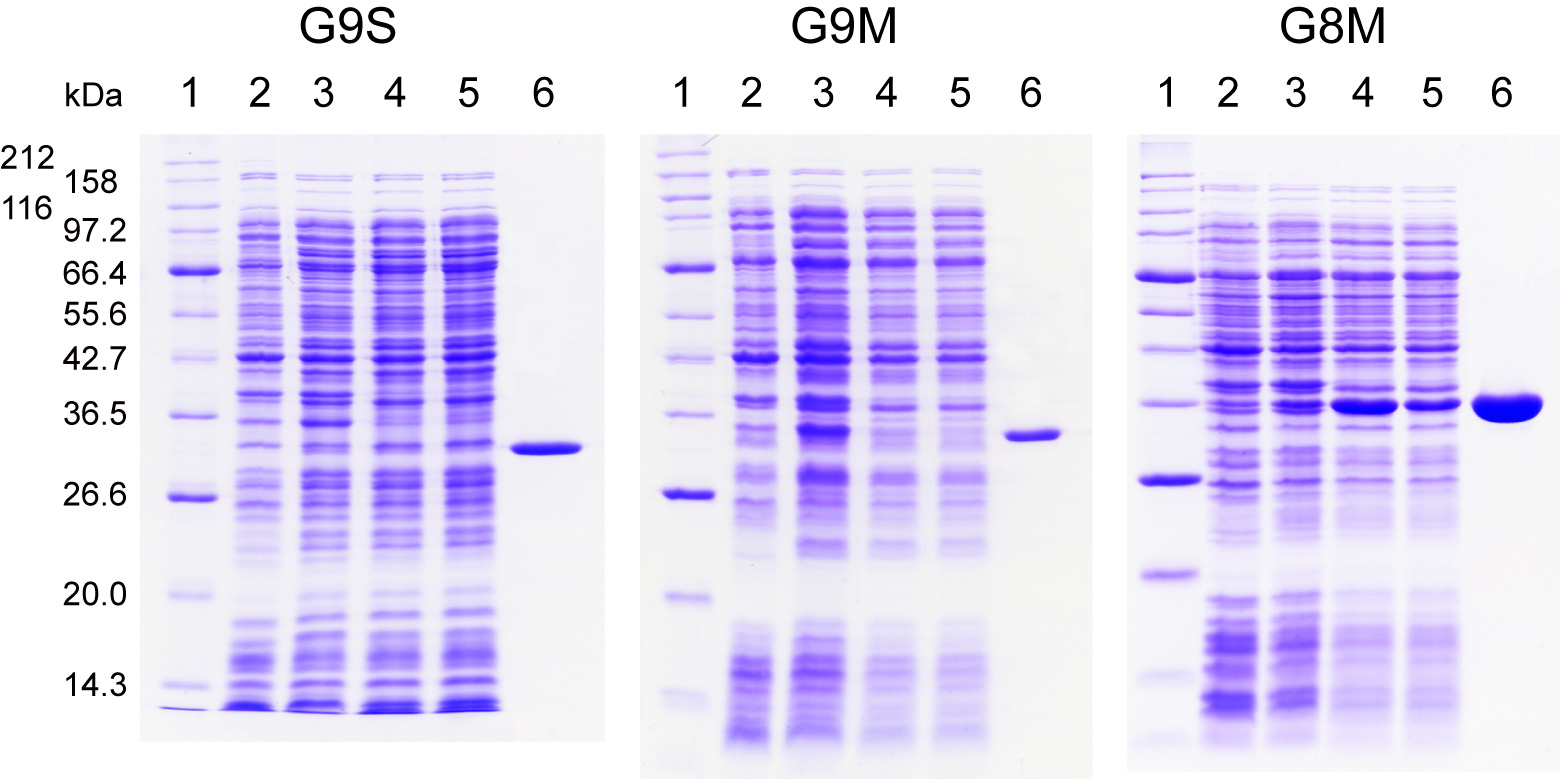
Fig. 1. Expression and purification profiles of human galectin-9S (G9S),
galectin-9M (G9M), and galectin-8M (G8M) on SDS-PAGE.
Samples at each expression and purification step for G9M and G8M were electrophoretically separated in SDS/12.6% polyacrylamide gels under reducing conditions, and then stained with Coomassie brilliant blue R-250. Lane 1, molecular weight marker proteins; lane 2, E. coli lysate (uninduced control); 3, E. coli lysate (16 h after induction); 4, E. coli extract; 5, non-adsorbed fraction from lactose-agarose; 6, lactose eluate from lactose-agarose.

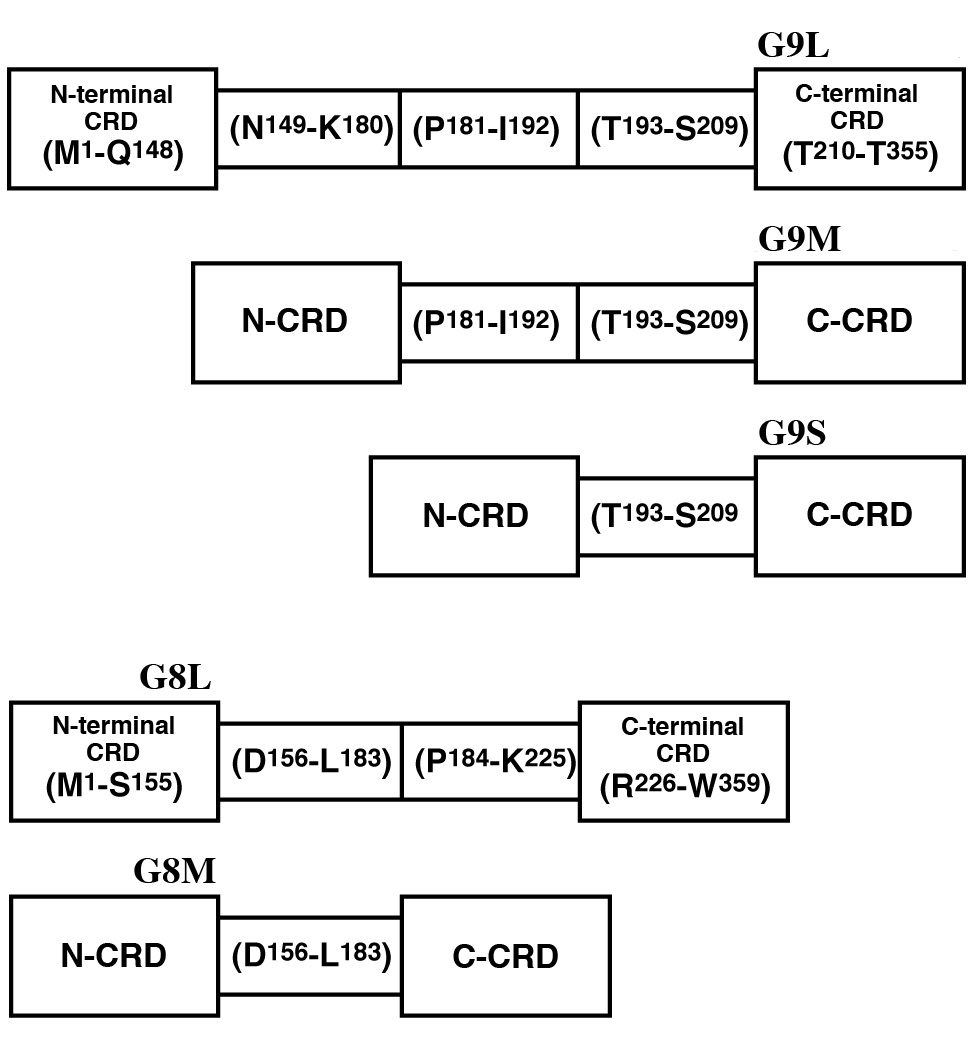
Fig. 2. Schematic representation of human galectin-9 and galectin-8 isoforms.
Superscript numbers indicate amino acid positions in galectin-9L and galectin-8L. The N- and C-terminal CRDs (N-CRD and C-CRD), and the linker peptide region were tentatively assigned based on intron-exon structures. However, X-ray crystallographic studies and phylogenetic analyses predict that Val15-Gln148 (Ile17-Ser155) and Met225-Thr355 (Leu227-Trp359) form the stably folded cores of N-CRD and C-CRD of galectin-9L (galectin-8L), respectively. |
| Copyrights |
 Attribution-Non-Commercial Share Alike Attribution-Non-Commercial Share Alike
This work is released underCreative Commons licenses
|
| Date of registration:2014-07-29 16:59:26 |
- Sato, M., Nishi, N., Shoji, H., Seki, M., Hashidate, T., Hirabayashi, J., Kasai, K., Hata, Y., Suzuki, S., Hirashima, M., and Nakamura, T. (2002) Functional analysis of the carbohydrate recognition domains and a linker peptide of galectin-9 as to eosinophil chemoattractant activity. Glycobiology 12, 191–197 [PMID : 11971863]
- Hirashima, M., Kashio, Y., Nishi, N., Yamauchi, A., Imaizumi, T.A., Kageshita, T., Saita, N., and Nakamura. T. (2004) Galectin-9 in physiological and pathological conditions. Glycoconj J. 19, 593–600 [PMID : 14758084]
- Nishi, N., Itoh, A., Fujiyama, A., Yoshida, N., Araya, S., Hirashima, M., Shoji, H., and Nakamura, T. (2005) Development of highly stable galectins: truncation of the linker peptide confers protease-resistance on tandem-repeat type galectins. FEBS Lett. 579, 2058–2064 [PMID : 15811318]
- Itoh, A., Fukata Y., Miyanaka, H., Nonaka, Y., Ogawa, T., Nakamura, T., and Nishi, N. (2013) Optimization of the inter-domain structure of galectin-9 for recombinant production. Glycobiology 23, 920–925 [PMID : 23507964]
|
This work is licensed under Creative Commons Attribution-Non-Commercial Share Alike. Please include the following citation
How to Cite this Work in an article:
Nishi, Nozomu,
Nakamura, Takanori,
(2014). GlycoPOD https://jcggdb.jp/GlycoPOD.
Web.25,4,2024 .
How to Cite this Work in Website:
Nishi, Nozomu,
Nakamura, Takanori,
(2014).
Expression and purification of recombinant human galectin-9.
Retrieved 25,4,2024 ,
from https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t105.
html source
Nishi, Nozomu,
Nakamura, Takanori,
(2014).
<b>Expression and purification of recombinant human galectin-9</b>.
Retrieved 4 25,2024 ,
from <a href="https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t105" target="_blank">https://jcggdb.jp/GlycoPOD/protocolShow.action?nodeId=t105</a>.
Including references that appeared in the References tab in your work is
much appreciated.
For those who wish to reuse the figures/tables, please contact JCGGDB
management office (jcggdb-ml@aist.go.jp).
|
|